Mass spectrometry technique uncovers an elusive sugar modification in stem cells
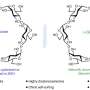
Recently, O-linked N-acetylglucosamine (O-GlcNAc), a sugar ring that reversibly modifies proteins inside the cell nucleus and cytoplasm in a process known as O-GlcNAcylation, has been revealed to be a key regulator of cell signaling in the body. Researchers believe that cellular changes induced by O-GlcNAc may be linked to chronic diseases, such as diabetes and cancer. Characterizing such dynamic relationships, however, is challenging.
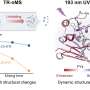
Normally, O-GlcNAc detection relies on 'shotgun' mass spectrometry (MS), which breaks proteins into sequence-specific peptide chains and measures their molecular weight. But as O-GlcNAcylated peptides are present in such small amounts, building up concentrations sufficient for detection requires time-consuming and tedious enrichment methods.
Now, Julien Maury and Andre Choo from the A*STAR Bioprocessing Technology Institute in Singapore and co-workers have developed a simple way to detect native O-GlcNAcylated proteins—even at 10,000-fold dilution—with a targeted MS technique known as multiple reaction monitoring (MRM).
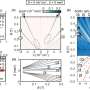
Instead of scrutinizing all possible peptides from a protein system, MRM-MS filters out masses that correspond to an expected precursor ion. Then, the joined-up precursor peptide and selected fragment ions—termed 'transition couples'—are monitored to build up a quantitative assay of the protein structure.
Choo explains that by focusing on specific O-GlcNAc-modified targets, MRM-MS enhances detection limits without extensive labeling or enrichment techniques. Furthermore, he notes that the method's ability to spot sugar modifications in complex peptide mixtures could greatly simplify cell bioanalysis.
After programming the MRM-MS system to spot standard O-GlcNAcylated peptides, the team turned their attention to the enzyme glycogen synthase kinase-3 beta (GSK-3β), which competes with O-GlcNAc for binding sites on proteins and is linked to numerous high-profile diseases. Although unconfirmed, GSK-3β may itself be modified and regulated by O-GlcNAc.
To investigate O-GlcNAcylated GSK-3β peptides, the researchers used gel electrophoresis to extract GSK-3β from proteins derived from human embryonic stem cells (hESCs). Subsequent MRM-MS analysis of the sample revealed the presence of a novel O-GlcNAcylated GSK-3β peptide with three potential binding sites—a modification with potential significance for the enzyme's autoinhibition mechanism. Using their technique, the researchers could also quantify changes in O-GlcNAcylated GSK-3β peptide levels following hESC drug treatment.
Choo and Maury anticipate that their work could help to detect and quantify O-GlcNAcylated peptides in samples where protein amounts are scarce, such as mouse brains. "Additionally, scientists want to know the dynamic variations in O-GlcNAc following drug treatment and cell differentiation studies," they say. "Quantification by MRM-MS is exactly suited to these sort of investigations."
More information: Maury, J. J. P., Ng, D., Bi, X., Bardor, M. & Choo, A. B.-H. "Multiple reaction monitoring mass spectrometry for the discovery and quantification of O-GlcNAc-modified proteins." Analytical Chemistry 86, 395–402 (2014). dx.doi.org/10.1021/ac401821d
Journal information: Analytical Chemistry