How do bacteria repair damage from the sun?
(Phys.org) —From bacteria to plants to humans, all organisms have mechanisms that they use to repair DNA damaged by ultraviolet (UV) light. This fundamental maintenance function is critical to our health because damaged DNA can lead to diseases such as cancer. So, of course, we know all about how it works. Or so scientists thought. New research has provided evidence that the current model for how UV repair functions must to be reworked.
At issue is the number of molecules of the important proteins in the complex, UvrA and UvrB. At issue is the number of molecules of the important proteins in the complex, UvrA and UvrB. This new study, conducted by researchers from Harvard University utilizing the U.S. Department of Energy Office of Science's Advanced Photon Source promises to provide new insights into the fundamental mechanisms of DNA repair and into diseases that are caused by mutations in these genes such as xeroderma pigmentosum (an autosomal recessive genetic disorder of DNA repair in which the ability to repair damage caused by ultraviolet (UV) light is deficient), Cockayne syndrome (a disorder characterized by short stature and an appearance of premature aging), and trichothiodystrophy (an inherited condition in which hair is brittle, sparse, and easily broken).
Previous work had shown that the bacterial DNA repair mechanism involved three proteins, UvrA, UvrB, and UvrC. The model was that two copies of UvrA traveled around with one copy of UvrB (A2B1) and scanned the DNA for places where damage had occurred. When a spot was identified that needed repair, the A2B1 stopped and waited at the spot that needed repair. At this point UvrC would come in and displace UvrA and form a complex with UvrB (B1C1) that conducted the repair. This model explained how the complex could scan both DNA strands but then orient to repair just one strand. The asymmetric A2B1 fit perfectly.
The only problem was, data from various structural methods started to pile up that were not consistent with this model and suggested that the initial complex might actually be A2B2. In addition, the Harvard research team solved the crystal structure of the UvrA/UvrB complex and it was consistent with the A2B2 model. They proposed that A2B2 identified the damage and then the UvrA and one of the UvrB molecules were released when UvrC arrived to repair the problem. However, outstanding questions remained about whether there might be other configurations in the crystal structure consistent with the longstanding model.
The group decided the best way to answer these questions was to evaluate the complex using small-angle x-ray scattering (SAXS). Their thinking was that low-resolution structural data collected on the protein complex in solution would be ideal for comparison modeling of various protein configurations. In addition, they hoped to find evidence for structural changes associated with how these proteins bind to ATP and use its energy for their activities.
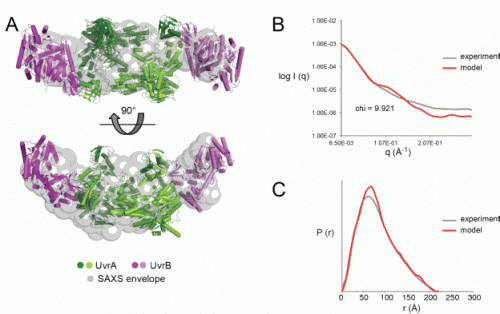
SAXS analysis of a solution of UvrA and UvrB in complex carried out at the Biophysics Collaborative Access Team (Bio-CAT) beamline 18-ID-D at the Argonne Advanced Photon Source showed an elongated structure (see the figure).
Modeling of the SAXS data against five possible configurations, four A2B2 options and one A2B1 option, showed that the data was consistent with one of the A2B2 configurations observed in the crystal structure but ruled out the others. Further analysis of the SAXS data by five other methods also supported this conclusion.
Unfortunately, when the group added ATP or ADP to the complex, they were not able to see any significant changes. This may require further experiments with different versions of the protein that allow them to keep the proteins bound to either ATP or ADP. For now, the Harvard researchers will be working on their new model to explain how bacteria perform this universal function.
More information: Danaya Pakotiprapha, David Jeruzalmi. "Small-angle X-ray scattering reveals architecture and A2B2 stoichiometry of the UvrA–UvrB DNA damage sensor," Proteins 81,132 (2013). DOI: 10.1002/prot.24170
Provided by Argonne National Laboratory