Soil pretreatment boosts protein recovery for microbial community studies
Scientists have significantly boosted the recovery and identification of proteins expressed by soil-dwelling microbes over what was previously possible, thanks to a new method of soil pretreatment being used at EMSL in Richland, Wash. The new strategy for processing samples reveals additional insight into the function of microbial communities in their native environments.
Studying the proteins expressed by soil-dwelling microbial communities helps to define their fundamental biogeochemical roles in carbon cycling, nitrogen cycling, phosphorus cycling, and climate regulation. It also helps to determine how microbial communities might assist with environmental cleanup—for example, bacterial protein expression could be used to remediate sites contaminated with toxic metals or radionuclides.
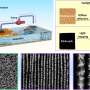
Metaproteomics studies, or the characterization of all proteins in a cellular community, are imperative for understanding the function of microbial communities that live in soil—but it is sticky business. Methods for protein recovery and identification that rely on simply lysing microbes in place results in the proteins binding irretrievably to the soil, and attempts to separate bacteria from soil prior to lysis are incomplete at best. To address this issue, PNNL staff tested a variety of methods and found that recovery and identification of proteins is significantly enhanced if, before lysis, soil samples are treated with both a cocktail of polar positive amino acids, which bind to soil and block protein binding sites, and desorption buffer, which enhances the release of proteins from the soil/solution mixture. This method proved effective in a model system consisting of E. coli and uncontaminated soil samples from Rifle, Colorado; a variety of soil samples having different sand and silt compositions; and an indigenous soil sample contaminated with diesel oil from King George, Antarctica. The method does not interfere with mass spectrometry analyses, and in the model system, the research team showed that the polar positive amino acid/desorption buffer pretreatment improved protein identification by nearly an order of magnitude compared to using no pretreatment at all.
More information: Nicora, C. et al. 2013. Amino acid treatment enhances protein recovery from sediment and soils for metaproteomic studies, Proteomics. DOI: 10.1002/pmic.201300003
Journal information: Proteomics
Provided by Environmental Molecular Sciences Laboratory