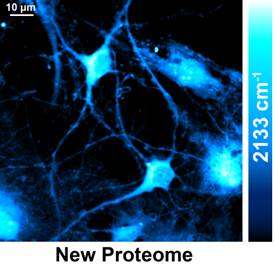
Watching the production of new proteins in live cells
Researchers at Columbia University, in collaboration with biologists in Baylor College of Medicine, have made a significant step in understanding and imaging protein synthesis, pinpointing exactly where and when cells produce new proteins. Assistant Professor Wei Min's team developed a new technique to produce high-resolution imaging of newly synthesized proteins inside living cells. The findings were published in the July 9th issue of The Proceedings of the National Academy of Sciences (Volume 110; Issue 28).
Proteins carry out almost every crucial biological function. Synthesis of new proteins is a key step in gene expression and is a major process by which cells respond rapidly to environmental cues in physiological and pathological conditions, such as cancer, autism and oxidative stress. A cell's proteome (i.e., the sum of all the cell's proteins) is highly dynamic and tightly regulated by both protein synthesis and disposal to maintain homeostasis and ensure normal functioning of the body. Many intricate biological processes, such as cell growth, differentiation and diseases, involve new protein synthesis at a specific location and time. In particular, long-lasting neuronal plasticity (changes in neural pathways and synapses that come from alterations in behavior, environment and bodily injury), such as those underlying learning and long-term memory, require new protein synthesis in a site- and time- dependent manner inside neurons.
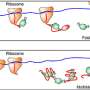
Min and colleagues' new technique harnesses deuterium (a heavier cousin of the normal hydrogen atom), which was first discovered by Harold Urey in 1932, also at Columbia University. When hydrogen is replaced by deuterium, the biochemical activities of amino acids change very little. When added to growth media for culturing cells, these deuterium-labeled amino acids are incorporated by the natural cell machineries as the necessary building blocks for new protein production. Hence, only newly synthesized proteins by living cells will carry the special deuterium atoms connected to carbon atoms. The carbon-deuterium bonds vibrate at a distinct frequency, different from almost all natural chemical bonds existing inside cells.
The Columbia team utilized an emerging technique called stimulated Raman scattering (SRS) microscopy to target the unique vibrational motion of carbon-deuterium bonds carried by the newly synthesized proteins. They found that by quickly scanning a focused laser spot across the sample, point-by-point, SRS microscopy is capable of delivering location-dependent concentration maps of carbon-deuterium bonds inside living cells.
"Incorporation of deuterium-labeled amino acids to new proteins is minimally disruptive, and their biochemical properties are almost identical to their natural counterparts," says Lu Wei, the lead author of the paper. "Our technique is highly sensitive, specific, and compatible with living systems under physiological conditions that don't require killing cells or staining."
Prior to this discovery, the ability to observe protein synthesis in living cells had eluded scientists, who devoted extensive efforts to achieving this goal. A classic strategy that involves labeling amino acids with radioisotopes to trace and quantify proteome dynamics requires the samples be killed and exposed to films. Fluorescence microscopy, another popular method, takes advantage of the inherent glowing of green fluorescent protein (GFP) to follow a protein. While this process does work on individual proteins, scientists can't observe the cell's entire proteome. A third technique, bioorthogonal noncanonical amino acid tagging (BONCAT) metabolically incorporates unnatural (biosynthetic) amino acids containing reactive chemical groups. However, the method generally requires killing cells and subsequent dye staining, a process that presents an issue for live tissues and animals. Therefore, it is extremely challenging and desirable to quantitatively image proteome synthesis in living cells, tissues and animals with high resolution. Min's research opens the door for a new method to study living cells, presenting opportunities to understand previously unanswered questions about the behavior of cells as they perform their functions.
The next step for Min's team is to capture where and when a new protein is produced inside brain tissues when an animal is subject to various lab exercises to form long-term memory. "Our new technique will greatly facilitate understanding the molecular mechanisms of many complex behaviors such as learning and diseases," he says.
Journal information: Proceedings of the National Academy of Sciences
Provided by Columbia University