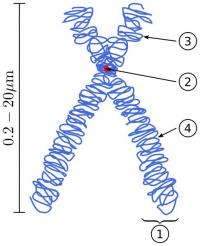
Scheme of a Chromosome. (1) Chromatid. One of the two identical parts of the chromosome after S phase. (2) Centromere. The point where the two chromatids touch, and where the microtubules attach. (3) Short arm (4) Long arm. In accordance with the display rules in Cytogenetics, the short arm is on top. Image: Wikipedia.
(Phys.org) -- Fruit flies are commonly used in genetics research because their lifespan is short, they are easy to breed in the laboratory, and mutants are widely available. There are about 1,500 known species. Now a new study of one of these species of fruit fly has tracked the evolution of a pair of sex chromosomes that appeared only around a million years ago.
The X and Y chromosomes in the fruit fly are, like the human X and Y chromosomes, vastly different in size and base sequence. In humans the chromosome pair is thought to have begun to evolve about 200 million years ago from an original closely matched pair of autosomal (non-sex) chromosomes. At present the Y chromosome contains only around 50 genes, while the much larger X chromosome contains about 1,000. In most species the evolution from autosome to sex chromosome occurred so long ago their evolution is difficult to track because there are few remnants of their origins.
The research team, Assistant Professor Doris Bachtrog and Dr. Qi Zhou of the Department of Integrative Biology at the University of California, Berkeley, decided to try to shed some light on sex chromosome evolution by studying the genome of Drosophila miranda flies, in which “neo-X” and “neo-Y” chromosomes first appeared when the Y chromosome fused with an autosome only about one million years ago. In a related species, Drosophila pseudoobscura, no fusion occurred, and so by comparing the genomes of the two species, the researchers were able to demonstrate how the X and Y chromosomes have evolved in D. miranda.
According to Bachtrog, when the neo-X and neo-Y chromosomes formed, about 3,000 genes were sex-linked, with female flies having two copies of neo-X and male flies having one copy each of neo-Y and neo-X. In the million years since the fusion, the Y chromosome shows massive degeneration, with over a third of the neo-Y genes having lost their function. The degeneration was already known to occur, but Prof. Bachtrog said the speed at which it had occurred was surprising.
Along with genes that have lost their function, other genes on the Y chromosome have evolved to be beneficial to males and expressed in specifically male tissues such as the prostate gland and testes. A similar evolution is also occurring on the X chromosome, on which genes expressed in female tissues are becoming more dominant.
The process of the genes becoming more beneficial to females is thought to take longer on the X chromosome because males also contain a copy of the X, leading to a slower distribution of these genes than in the Y, which is only found in males. The evolution of the X chromosome is not only slower, but includes larger events, such as incorporation of genes from other chromosomes into the X.
Bachtrog pointed out that in fruit flies some sex chromosomes have reverted to autosomes, and it is also possible that in Drosophila species the Y chromosome could eventually disappear altogether, and another mechanism for determining sex could then evolve.
More information: Sex-Specific Adaptation Drives Early Sex Chromosome Evolution in Drosophila, Science 20 July 2012: Vol. 337 no. 6092 pp. 341-345. DOI: 10.1126/science.1225385
ABSTRACT
Most species’ sex chromosomes are derived from ancient autosomes and show few signatures of their origins. We studied the sex chromosomes of Drosophila miranda, where a neo-Y chromosome originated only approximately 1 million years ago. Whole-genome and transcriptome analysis reveals massive degeneration of the neo-Y, that male-beneficial genes on the neo-Y are more likely to undergo accelerated protein evolution, and that neo-Y genes evolve biased expression toward male-specific tissues—the shrinking gene content of the neo-Y becomes masculinized. In contrast, although older X chromosomes show a paucity of genes expressed in male tissues, neo-X genes highly expressed in male-specific tissues undergo increased rates of protein evolution if haploid in males. Thus, the response to sex-specific selection can shift at different stages of X differentiation, resulting in masculinization or demasculinization of the X-chromosomal gene content.
Journal information: Science
© 2012 Phys.org