3D structure opens new avenue for drug discovery
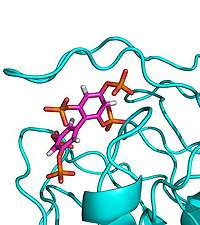
(PhysOrg.com) -- An international consortium has determined the structure of an important new drug target in complex with a synthetic molecule designed by our researchers, opening up new avenues for drug discovery.
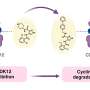
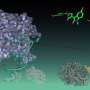
The enzyme SHIP2, which plays a major role in cell signalling, has attracted particular attention due to its role in the negative regulation of insulin signalling and is thought to be involved in type-2 diabetes, obesity and cancer. Previous attempts to crystallise the enzyme with its natural substrate were unsuccessful, so scientists at Bath designed a new synthetic inhibitor as a mimic and with their European colleagues solved the X-ray crystal structure of a key fragment of SHIP2 bound to this compound.
The research, recently published in the leading international journal ACS Chemical Biology, was a collaborative project carried out by an international group of scientists based at the Karolinska Institutet in Stockholm Sweden, Nanyang University Singapore, the Université Libre de Bruxelles in Belgium and the University of Bath.
The scientists also undertook computational molecular dynamics on the complex, and discovered a flexible loop region of the protein that may close over the compound during binding. The researchers hope that targeting such a closed complex could provide a new strategy for the design of small-molecule drugs against SHIP2.
Professor Barry Potter, who led the enterprise together with his Wellcome Trust–funded Bath colleagues Drs Steve Mills, Andrew Riley, Gyles Cozier and Mark Thomas, said: “Such interdisciplinary collaboration represents a real route to early progress in drug discovery at a time when the global pharmaceutical industry is restructuring and looking more than ever towards academic-industry partnerships for early stage drug discovery, rather than in-house R&D.
“These data further reinforce use of a new class of synthetic molecule that we have pioneered at Bath for several years, for co-crystallisation studies.
“This work emphasises the strength of Medicinal Chemistry at the University of Bath and demonstrates that academic scientists can play a key role in drug discovery, particularly at early and innovative stages.”
The next step will be to design in silico related, but more drug-like, compounds that might bind to the closed complex of the SHIP2 enzyme. The researchers hope that others will use their work as a starting point to design such novel drug candidates.
More information: pubs.acs.org/doi/pdf/10.1021/cb200494d
Provided by University of Bath