New study explores proteins in Yellowstone bacteria for biofuel inspiration
Studies of bacteria first found in Yellowstone's hot springs are furthering efforts at the Department of Energy's BioEnergy Science Center toward commercially viable ethanol production from crops such as switchgrass.
The current production of ethanol relies on the use of expensive enzymes that break down complex plant materials to yield sugars that are fermented into ethanol. One suggested cheaper alternative is consolidated bioprocessing, a streamlined process that uses microorganisms to break down the resistant biomass.
"Consolidated bioprocessing is like a one-pot mix," said Oak Ridge National Laboratory's Richard Giannone, coauthor on a BESC proteomics study that looked at one microorganism candidate. "You want to throw plant material into a pot with the microorganism and allow it to degrade the material and produce ethanol at the same time."
The BESC study focused on Caldicellulosiruptor obsidiansis, a naturally occurring bacterium discovered by BESC scientists in a Yellowstone National Park hot spring. The microorganism, which thrives at extremely high temperatures, breaks down organic material such as sticks and leaves in its natural environment, and scientists hope to transfer this capability to biofuel production tanks.
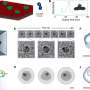
In a paper featured on the cover of the Journal of Proteome Research, the BESC team conducted a comparative analysis of proteins from C. obsidiansis grown on four different carbon sources, ranging from a simple sugar to more complex substrates such as pure cellulose and finally to switchgrass. The succession of carbon substrates allowed researchers to compare how the organism processes increasingly complex materials.
"This progression helps us look at how proteins change given different carbon substrates," Giannone said. "One of the goals is to identify new proteins that we haven't seen before. If an unknown protein doesn't show up on the simple sugars or cellulose, but it shows up on the switchgrass, then we can say something about that gene or protein—that it responds to something the switchgrass is providing."
The researchers found that growth on switchgrass prompted the organism to express an expanded set of proteins that deal specifically with the hemicellulose content of the plant, including hemicellulose-targeted glycosidases and extracellular solute-binding proteins. Acting together, these two sub-systems work to break down the plant material and import the resulting sugars into the cell. The scientists went on to show that once inside the cell, the organism "switches on" certain enzymes involved in pentose metabolism in order to further process these hemicellulose-derived sugars into usable energy.
"By comparing how C. obsidiansis reacted to switchgrass, relative to pure cellulose, we were able to pinpoint the specific proteins and enzymes that are important to plant cell wall deconstruction—a major roadblock to the production of advanced biofuels," Giannone said.
The team's measurement of the full complement and abundance of C. obsidiansis proteins, called proteomics, can now be combined with other technologies such as genomics, transcriptomics and metabolomics in order to obtain a 360-degree, system-wide view of the organism. Instead of studying discrete proteins, these systems biology-type approaches provide more thorough insight into the day-to-day operations of bioenergy-relevant organisms and thus better equip researchers to make recommendations about their use in ethanol production.
"One goal for us at the BioEnergy Science Center is to take these 'omic' technologies and integrate the data so we can draw definitive conclusions about a system," Giannone said.
More information: Coauthors on the paper are Hamburg University of Technology's Adriane Lochner and Garabed Antranikian, and ORNL's Martin Keller, David Graham and Robert Hettich. The full publication is available here: pubs.acs.org/doi/abs/10.1021/pr200536j
Provided by Oak Ridge National Laboratory