New optogenetic tools for biomedical research developed by UW scientists

(PhysOrg.com) -- Researchers at the University of Wyoming have characterized and engineered new proteins that expand the use of light as a tool to manipulate cell cultures, tissues and laboratory model animals.
In a paper published in the Journal of Biological Chemistry, UW researchers Min-Hyung Ryu, Oleg Moskvin, Jessica Siltberg-Liberles and Mark Gomelsky describe novel light-activated proteins to study cellular regulatory networks. They say this technology, called optogenetics, is beginning to revolutionize biomedical research. Optogenetic approaches have already been applied to investigating neural circuits relevant to locomotion and awakening, as well as to brain disorders such as Parkinson's disease, schizophrenia and epilepsy.
Animals have very few proteins that are naturally sensitive to light, says Siltberg-Liberles, director of the INBRE (IDeA Networks of Biomedical Research Excellence) Bioinformatics service core. She says optogenetics allows one to "borrow" natural light-activated proteins from microbes or plants and deliver their genes into model animal organisms.
"We are used to regulating cellular processes with drugs. However, it is difficult, if not impossible, to deliver drugs to specific cells and organs that need to be treated while sparing the rest of the organism from the unwanted side effects," says Gomelsky, an associate professor in the UW Department of Molecular Biology.
"Then, once a drug is administered, it takes time for it to act and to be removed from an organism, so we have relatively little control over timing," Gomelsky adds. "The beauty of optogenetic approaches is that light can be delivered by lasers with extremely high spatial precision; therefore, one can manipulate only target cells. Furthermore, turning light on and off takes a split second, so one gains high temporal precision as well."
"It would be even more powerful if we could design proteins that have desired functions to be turned on by light of a specific waveband," Siltberg-Liberles says. "However, at present, designing light-activated proteins is only half-science and half-art."
There is a high demand for developing better light switches and a new set of functions that can be controlled by light, says Siltberg-Liberles. She explains that two molecules, cAMP and cGMP, control a variety of cellular processes, such as cell growth, blood glucose levels, cardiac function, learning, memory, cancer cell survival and others. These two molecules are synthesized by the enzymes called adenylyl and guanylyl cyclases. The UW team worked on making the light-activated versions of these cyclases.
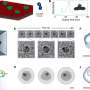
"We have identified a Blue-Light activated Adenylyl Cyclase, BlaC, in a genome of the marine bacterium Beggiatoa. After characterizing the protein, we found that it is better controlled by light than any protein of that function known to date," she says. "Using computational and protein engineering approaches, we redesigned BlaC to function as a light-activated guanylyl cyclase, a new activity that has never previously existed."
Siltberg-Liberles says as soon as the paper appeared in the online early version of Journal of Biological Chemistry, the researchers started receiving requests by research groups in the United States and Europe that wanted to use the light-activated cyclases to study diverse processes in various animal systems.
"Our paper has been recommended by a Faculty of 1000 Biology, a group of top experts that highlights noteworthy research findings," she says.
Research on light-activated proteins continues. UW scientists are now engineering infrared-light activated cyclases, which Gomelsky says will be important for biomedical research because infrared light penetrates animal tissues much deeper than blue light.
"We are grateful to INBRE for partial support of our light-activated protein engineering project and also for creating the Bioinformatics service core. Without access to Jessica's expertise, this project would not have been possible," says Gomelsky.
Jun Ren, INBRE director, says the Bioinformatics service core was established to add computational resource and expertise in bioinformatics to experimental life scientists across campus and this is a good example of what it can do. Siltberg-Liberles adds that the Bioinformatics service core is available for all life scientists and new projects are always welcome.
More information: The entire article can be found at www.jbc.org/content/285/53/415 … e5-a8e4-512a00fe8ad7 .
Provided by University of Wyoming