Working in concert
As cells go about the process of building new proteins, each amino acid gets delivered individually to the protein-synthesizing machinery of the ribosome by a specialized transfer RNA (tRNA) molecule. This process is generally facilitated by a variety of RNA synthetase enzymes, with each type of amino acid-attached tRNA formed as the product of a distinct synthetase.
However, archaea and most bacteria lack the capacity to directly synthesize every necessary tRNA-amino acid conjugate. For example, the glutamine-linked tRNA (Gln-tRNAGln) in such species is instead produced via a two-step process, a synthetic pathway that is initiated by glutamyl-tRNA synthetase (GluRS), the enzyme normally responsible for producing the glutamate-bearing tRNA (Glu-tRNAGlu).
In this case, however, GluRS needs to make a ‘wrong move’ in order to produce an unusual hybrid tRNA, Glu-tRNAGln, which serves as an essential intermediate in the Gln-tRNAGln production process. This pathway has proven baffling to biochemists such as Shigeyuki Yokoyama, director of the RIKEN Systems and Structural Biology Center (SSBC) in Yokohama. “This mechanism has the possibility of forming harmful byproducts, including Gln-tRNAGlu and Glu-tRNAGln, which can cause serious errors in protein synthesis,” he says. “We wanted to know how such a risky system functions efficiently.”
To further add to the mysteries surrounding Gln-tRNAGln production, the complex responsible for the second step in this process, GatCAB, has been proposed to interact with the same region of Glu-tRNAGln as GluRS. This has left it unclear how these two enzymes are able to effectively partner with their common substrate in a manner that enables the two stages of synthesis to flow seamlessly without interfering with one another or potentially releasing toxic intermediates into the cellular environment.
Taking turns
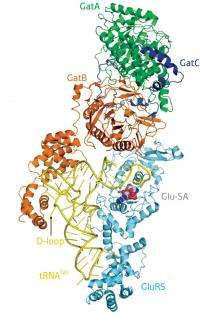
Yokoyama recently teamed up with SSBC colleague Takuhiro Ito in an effort to tackle these various mysteries, collecting high-resolution structural data from the bacterial species Thermotoga maritima for both the GluRS-tRNAGln complex as well as the ‘glutamine transamidosome’—the ternary complex containing tRNAGln, GluRS and GatCAB. They confirmed that the latter is indeed a stable complex (Fig. 1), indicating that some mechanism must exist that prevents the two enzymes from simultaneously occupying the same space and disrupting each other’s function. Yokoyama and Ito have published their findings in the science journal Nature.
Comparison of the two structures revealed that tRNAGln undergoes structural changes when assembled within the glutamine transamidosome, interacting with GatCAB via structural elements that are absent in tRNAGlu. In order to confirm this mechanism, Yokoyama and Ito constructed a series of hybrid tRNAGlu molecules that contain various elements from tRNAGln and examined the extent to which they could successfully be processed to yield Gln-tRNAGlu. Based on these experiments, they were able to confirm that the region in question—also known as the D loop—is an essential structural feature in this process, revealing the mechanism by which this complex avoids introducing inappropriate modifications to Glu-tRNAGlu. “Our structure clearly shows how GatCAB recognizes tRNAGln without inhibiting the reaction by GluRS,” says Yokoyama.
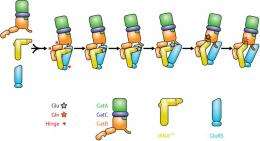
Their analysis also revealed a number of flexible ‘hinge’ segments in both GatCAB and GluRS that appear to contribute to their capacity to perform modifications in a stepwise manner (Fig. 2). Each of these factors initially binds to a different segment of tRNAGln, with the GatCAB interaction contributing the discrimination mechanism revealed by Yokoyama and Ito. GluRS subsequently transfers a glutamate amino acid to the tRNA and undergoes a structural rearrangement that frees the newly modified ‘acceptor arm’ domain of the tRNAGln molecule. GatCAB then executes a pivoting motion that brings its own catalytic domain in contact with the glutamate-bound acceptor arm, at which point it initiates the amidation chemical reaction necessary to transform this amino acid into a glutamine residue. “This structure itself is very unique and interesting,” says Yokoyama. “One substrate tRNA is bound by two different enzymes working consecutively.”
Protein evolution—past and future
Based on the unusual mechanism that they identified, the researchers hypothesize that this process of tRNA modification may have emerged relatively recently in the evolution of protein synthesis. As such, glutamine and asparagine—another amino acid whose synthesis is also the result of a two-step synthesis in a number of bacterial and archaeal species—could represent late additions to the genetic code.
In other work, Yokoyama and colleagues have explored methods for producing proteins whose characteristics have been selectively modified through the inclusion of atypical or synthetic amino acids, typically by inducing cells to express altered versions of tRNA synthetase enzymes that exclusively work with these alternative amino acids. However, the pathway described here could provide a useful model for future engineering efforts. “It might be possible to incorporate a new unnatural amino acid into the genetic code via this mechanism—modifying amino acids after tRNA formation, but before they are transferred to the ribosome,” explains Yokoyama.
However, all of the data presented in their study of Gln-tRNAGln represent deductions drawn from only a relatively small number of structural ‘snapshots’, based on a transamidosome complex that was essentially trapped in the first stage of Gln-tRNAGln synthesis. In order to further understand the detailed operation of this cellular machine, Yokoyama now intends to collect additional structural data that will help fill in the gaps and provide additional confirmation for the model that he and Ito have developed. “We have not succeeded yet, but we would like to continue our trials to crystallize the amidation form of the transamidosome,” he says.
More information: Ito, T. & Yokoyama, S. Two enzymes bound to one transfer RNA assume alternative conformations for consecutive reactions. Nature 467, 612–616 (2010).
Provided by RIKEN