Researchers find 19 million-year-old genomic fossils of hepatitis B-like viruses in songbirds
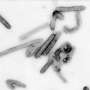
(PhysOrg.com) -- Biologists from The University of Texas at Arlington have uncovered virus fragments from the same family as the modern Hepatitis B virus locked inside the genomes of songbirds such as the modern-day zebra finch.
"And they've been sitting there for at least 19 million years, far longer than anyone previously thought this family of viruses had been in existence," say Cédric Feschotte and Clément Gilbert, co-authors of the new study being published today in PLoS Biology, the flagship journal of the Public Library of Science. Feschotte is an associate professor and member of the UT Arlington Genome Biology Group. Gilbert is a post-doctoral research associate in the group.
The article, entitled "Genomic Fossils Calibrate the Long-Term Evolution of Hepadnaviruses," marks the first time that endogenous hepadnaviruses have been found in any organism. An endogenous virus is one that deposits itself or fragments of itself into the chromosome of an organism, allowing it to be passed from generation-to-generation. Previously, most of these known "fossilized" virus sequences came from retroviruses.
Feschotte and Gilbert's results also suggest that the birds could be carriers of these types of viruses today.
Gilbert and Feschotte dated the hepadnavirus fragments by locating them in the same spot on the genome of five species of passerine birds and then tracing those species to a common ancestor that lived more than 19 million years ago.
Eddie Holmes, a distinguished professor of biology at Penn State University's Eberly College of Science and an expert in the field of viral evolution, said Feschotte and Gilbert's work "provides a glimpse into an ancient viral world that we never knew existed."
"The results they obtained were remarkable; whereas we previously thought of hepadnavirus evolution on time-scales of only a few thousand years, this paper shows that the true time-scale is in fact many million years. Therefore, hepadnavirues, and likely many other viruses as well, are far older than we previously thought." Holmes said.
In another surprising finding, the older versions of the hepadnaviruses are remarkably similar to today's viruses. Feschotte believes that the slow evolution of the hepadnaviruses observed in birds indicates that the viruses are, in the long run, better adapted to their hosts than what is suggested by study of the disease-causing Hepatitis B viruses.
"Genomic fossils like the remarkable hepadnaviral fossils found by Gilbert and Feschotte have the prospect of completely revising our preconceived notions about the age and evolution of such viruses," said Harmit Singh Malik, an associate member of the Fred Hutchinson Cancer Research Center in Seattle and one of the leaders in the new field of ‘paleovirology'. "They provide an unexpectedly clear lens on an ancient time when these viruses were prevalent and abundant."
The study also opens avenues for research that might help predict and prevent human viral pandemics originating in bird species.
"Given that they were infected in the past, it is legitimate to think that some of these birds may still carry such viruses today," said Gilbert. "We can therefore use this discovery as a guide to screen targeted groups of bird species for the presence of new circulating Hepatitis B-like viruses. "
More information: Gilbert C, Feschotte C (2010) Genomic Fossils Calibrate the Long-Term Evolution of Hepadnaviruses. PLoS Biol 8(9): e1000495. doi:10.1371/journal.pbio.1000495
Provided by University of Texas at Arlington