Boon to plant science

In both plant and animal cells, protein activity is often regulated by phosphorylation, by which a phosphate group is added to one or more sites on a protein. A team led by Ken Shirasu of RIKEN Plant Science Center, Yokohama, Japan, has found very similar patterns of protein phosphorylation even in distantly related plant species, a discovery that should advance plant engineering. The data is now freely available online from RIKEN's new Plant Phosphoproteome Database.
Just as a 'genome' includes all of the hereditary information of an organism encoded in its DNA, the entire suite of phosphorylated proteins and their phosphorylation sites found within an organism constitutes its ‘phosphoproteome’. “Elucidating and comparing the phosphoproteomes of different species is key to understanding diverse biological phenomena, including growth and development,” explains Shirasu.
Until now, the only plants whose phosphoproteomes have been studied in any detail are model species such as Arabidopsis thaliana, which belongs to a group of flowering plants known as the dicots. To broaden knowledge of plant phosphoproteomes, Shirasu and his colleagues embarked on the first large-scale protein phosphorylation screen in rice (Oryza sativa). Unlike Arabidopsis, whose phosphoproteome they have also updated, rice is a monocot.
“Rice is an economically important crop species providing the staple diet of millions of people, and since its genome has been sequenced it has also become an important model plant species,” says Shirasu. “Moreover, because rice is a monocot, it was interesting for us to compare its phosphoproteome with that of Arabidopsis, which being a dicot is only distantly related to rice in evolutionary terms.”
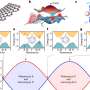
The researchers identified over 5,000 unique phosphorylation sites on a total of 3,393 proteins expressed in cultured rice cells. Of these, more than half are also phosphorylated in Arabidopsis, often sharing the same phosphorylation sites—despite their distant evolutionary relationship. Thus, the two species showed considerable overlap in the composition of phosphorylated proteins. The researchers also found significant similarities between the recently characterized phosphoproteome of Medicago truncatula, a small clover-like plant, with those of rice and Arabidopsis.
“What our findings show is that phosphorylation patterns are remarkably well conserved among plant species, meaning that what we learn from one species can be applied to another with relative ease,” says Shirasu.
The researchers hope that their findings will facilitate further comparative studies of plant phosphoproteomes, leading to better understanding of core regulatory mechanisms in plants, and the engineering of agronomically important plant species.
More information:
-- Nakagami, H., Sugiyama, N., Mochida, K., Daudi, A., Yoshida, Y., Toyoda, T., Tomita, M., Ishihama, Y. & Shirasu, K. Large-scale comparative phosphoproteomics identifies conserved phosphorylatinon sites in plants. Plant Physiology 153, 1161-1174 (2010). www.plantphysiol.org/cgi/conte … hort/pp.110.157347v1
-- RIKEN Plant Phosphoproteome Database phosphoproteome.psc.database.riken.jp/
Provided by RIKEN