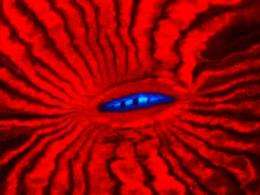
The image shows fluorescence of the mouth region of the reef coral Lobophyllia hemprichii. The red fluorescent protein isolated from this coral was rendered in an advanced tool for super-resolution microscopy. Credit: Jörg Wiedenmann
Scientists in Southampton, UK, and Ulm and Karlsruhe in Germany have shown that a variant form of a fluorescent protein (FP) originally isolated from a reef coral has excellent properties as a marker protein for super-resolution microscopy in live cells. Their findings have been published online by Nature Methods and will appear in print in the upcoming August issue of that journal.
Fluorescent proteins produced by a range of marine animals glow with a rainbow of colours, adding to the visual spectacle of coral reefs. Over recent years, molecular biologists have isolated a number of FPs and their genes, and used them to create genetically engineered variant FPs with particular light-emission characteristics.
"Fluorescent pigments from corals and related animals have proved to be invaluable lead structures to produce advanced markers for biomedical research," said Dr Jörg Wiedenmann of the University of Southampton's School of Ocean and Earth Science (SOES) based at the National Oceanography Centre, Southampton: "They enable a plethora of exciting experiments, including non-invasive study of dynamical processes within live cells,"
Photoactivatable FPs (PA-FPs)can, as their name suggests, be switched on by light. When light of a particular wavelength is shone upon them they start to glow, emitting light of characteristic hue.
Wiedenmann and his collaborators previously described EosFP, a PA-FP from the reef-building coral Lobophyllia. Genetic engineering yielded the variant IrisFP with dual photoactivation capacity. In one mode it is irreversibly 'photo-converted' from a green- to a red-emitting form under violet light. In a second mode, these two light-emitting forms can be switched on and off more or less at will using light of different wavelengths ('photo-switching').
For use in cell biology experiments, PA-FPs are genetically fused to proteins of interest, and expressed in live cells. Small regions of the cell are then illuminated with laser light of specific wavelength, causing the marker proteins to emit light at another wavelength. This allows dynamical cell processes to be visualised and studied under the microscope.
In the native state, four molecules of IrisFP join together to form a tetramer, creating problems for fusion-protein applications. To get round this, the researchers have now modified the protein by introducing four mutations. This makes individual IrisFP molecules (monomers) more stable, reducing their tendency for form tetramers.
"The monomeric variant, mIrisFP, maintains dual photoactivation capacity and has excellent properties as a genetically encoded fluorescent marker protein," explained Wiedenmann.
To test the usefulness of mIrisFP, the researchers genetically fused it with a number of other proteins within cultured cells. These included transcription factors, which regulate the expression of genes within the cell nucleus, and constituent proteins of the cell skeleton ('cytoskeleton'). In all cases, the fusion proteins functioned normally.
Further experiments demonstrated that mIrisFP fusion proteins could, as hoped, be used to study dynamical processes within live cells with a spatial resolution beyond the limits of conventional light microscopy. Specifically, the researchers successfully combined so-called pulse-chase experiments with photoactivation localisation microscopy (PALM) imaging to follow the movement of fluorescently marked fusion proteins over time and at very high spatial resolution.
"The dual photoactivation capability and the monomeric nature of mIrisFP should allow cell biologists to perform a wider range of experiments than possible using only conventional PA-FPs," said Wiedenmann.
"Marine animals such as corals and anemones are not only beautiful and important for ecosystem functioning, but also as source of fluorescent proteins of enormous value to biomedical research," he added.
More information: Publication: Fuchs, J., Boehme, S., Oswald, F., Hedde, P. N., Krause, M., Wiedenmann, J. & Nienhaus, G. U. A photoactivatable marker protein for pulse-chase imaging with superresolution. Nature Methods (Published online: 4 July 2010). doi:10.1038/nmeth.147
Provided by National Oceanography Centre, Southampton