Biochemists manipulate fruit flavor enzymes
Would you like a lemony watermelon? How about a strawberry-flavored banana? Biochemists at The University of Texas Medical School at Houston say the day may be coming when scientists will be able to fine tune enzymes responsible for flavors in fruits and vegetables. In addition, it could lead to environmentally-friendly pest control.
In the advance online publication of Nature on Aug. 20, UT Medical School Assistant Professor C.S. Raman, Ph.D., and his colleagues report that they were able to manipulate flavor enzymes found in a popular plant model, Arabidopsis thaliana, by genetic means. The enzymes—allene oxide synthase (AOS) and hydroperoxide lyase (HPL)—produce jasmonate (responsible for the unique scent of jasmine flowers) and green leaf volatiles (GLV) respectively. GLVs confer characteristic aromas to fruits and vegetables.
Green leaf volatiles and jasmonates emitted by plants also serve to ward off predators. "Mind you plants can't run away from bugs and other pests. They need to deal with them. One of the things they do is to release volatile substances into the air so as to attract predators of the bugs," Raman said.
"Genetic engineering/modification (GM) of green leaf volatile production holds significant potential towards formulating environmentally friendly pest-control strategies. It also has important implications for manipulating food flavor," said Raman, the senior author. "For example, the aroma of virgin olive oil stems from the volatiles synthesized by olives. By modifying the activity of enzymes that generate these substances, it may be possible to alter the flavor of the resulting oils."
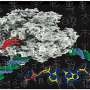
According to Raman, "Our work shows how you can convert one enzyme to another and, more importantly, provides the needed information for modifying the GLV production in plants." The scientists made 3-D images of the enzymes, which allowed them to make a small, but specific, genetic change in AOS, leading to the generation of HPL.
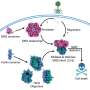
AOS and HPL are part of a super family of enzymes called cytochrome P450. P450 family enzymes are found in most bacteria and all known plants and animals. Although AOS or HPL are not found in humans, there are related P450 family members that help metabolize nearly half of the pharmaceuticals currently in use. In plants, AOS and HPL break down naturally-occurring, organic peroxides into GLV and jasmonate molecules. "Each flavor has a different chemical profile," Raman said.
"A notable strength of this manuscript is the combined use of structural and evolutionary biology to draw new insights regarding enzyme function. These insights led to the striking demonstration that a single amino acid substitution converts one enzyme into another, thereby showing how a single point mutation can contribute to the evolution of different biosynthetic pathways. This begins to answer the long-standing question as to how the same starting molecule can be converted into different products by enzymes that look strikingly similar," said Rodney E. Kellems, Ph.D., professor and chairman of the Department of Biochemistry & Molecular Biology at the UT Medical School at Houston.
The study dispels the earlier view that these flavor-producing enzymes are only found in plants, Raman said. "We have discovered that they are also present in marine animals, such as sea anemone and corals. However, we do not know what they do in these organisms."
Source: University of Texas at Houston