Researchers see history of life in the structure of transfer RNA
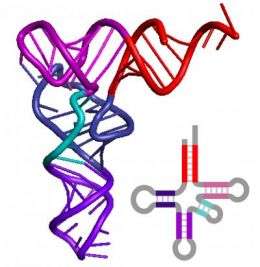
Transfer RNA is an ancient molecule, central to every task a cell performs and thus essential to all life. A new study from the University of Illinois indicates that it is also a great historian, preserving some of the earliest and most profound events of the evolutionary past in its structure.
The study, co-written by Gustavo Caetano-Anollés, a professor of crop sciences, and postdoctoral researcher Feng-Jie Sun, appears March 7 in PLoS Computational Biology. Caetano-Anollés is an affiliate of the U. of I. Institute for Genomic Biology.
Of the thousands of RNAs so far identified, transfer RNA (tRNA) is the most direct intermediary between genes and proteins. Like many other RNAs (ribonucleic acids), tRNA aids in translating genes into the chains of amino acids that make up proteins. With the help of a highly targeted enzyme, each tRNA molecule recognizes and latches onto a specific amino acid, which it carries into the protein-building machinery. In order to successfully add its amino acid to the end of a growing protein, tRNA must also accurately read a coded segment of messenger RNA, which gives instructions for the exact sequence of amino acids in the protein.
The fact that tRNA is so central to the task of building proteins probably means that it has been around for a long time, Caetano-Anollés said. His inquiry began with a hunch that understanding the structural properties of tRNA would shed light on how organisms and viruses evolved.
“Perhaps in evolution there are things that are so fundamental that they are kept, held onto, for millions or even billions of years,” Caetano-Anollés said. “Those are the fossils, the molecular fossils, that tell us about the past. Therefore, studying these molecules can address fundamental questions in biology and evolution.”
All tRNAs assemble themselves into a shape that, if flattened, resembles a cloverleaf. The team began by looking for patterns in this cloverleaf structure, using detailed data from hundreds of molecules representing viruses and each of the three superkingdoms of life: archaea, bacteria and eukarya.
The researchers converted all distinguishing features of the individual tRNA cloverleaf structures into coded characters, a process that allowed a computerized search for the most “parsimonious” (that is, the simplest, most probable) tRNA family tree. They conducted the same analysis on the tRNAs of each of the superkingdoms, to see how far these groupings diverged from the overall tree. This comparison allowed them to determine the order in which viruses and each of the superkingdoms diverged.
The new analysis supports an earlier study that suggested that the archaea were the first to arise as an evolutionarily distinguishable group. Archaea are microbes that can survive in boiling acid, near sulfurous ocean vents or in other extreme environments. The earlier study, also led by Caetano-Anollés, analyzed the vast catalog of protein folds – those precisely configured regions in proteins that give them their functionality – as a guidebook to evolutionary history.
“The transfer RNA data matches our earlier data,” Caetano-Anollés said. “This is important because two lines of independent evidence are supporting each other.”
The new analysis also indicates that viruses emerged not long after the archaea, with the superkingdoms eukarya and bacteria following much later – and in that order. This finding may influence the ongoing debate over whether viruses existed prior to, or after, the emergence of living cells, Caetano-Anollés said.
“This supports the idea that viruses arose from the cellular domain,” he said.
Source: University of Illinois at Urbana-Champaign