Astrotechnology Brings Nanoparticle Probes Into Sharper Focus
While pondering the challenges of distinguishing one nanosize probe image from another in a mass of hundreds or thousands of nanoprobes, two investigators at Emory University and the Georgia Institute of Technology made an interesting observation. The tiny, clustered dots of light looked a lot like a starry sky on a clear night.
The biomedical researchers realized that astronomers had already made great strides in solving a problem very similar to their own—isolating and analyzing one dot (in this case, a star) in a crowded field of light. They hypothesized that a computer system designed for stellar photometry, a branch of astronomy focused on measuring the brightness of stars, could hold the solution to their problem.
Now, May Wang, Ph.D., at Georgia Tech, and Shuming Nie, Ph.D., and their collaborators have created a technology based on stellar photometry software that provides more precise images of single molecules tagged with nanoprobes, particles specially designed to bind with a certain type of cell or molecule and illuminate when the target is found. The clearer images allow researchers to collect more detailed information about a single molecule, such as how the molecule is binding in a gene sequence, taking scientists a few steps closer to truly personalized and predictive medicine as well as more complex biomolecular structural mapping. In addition to biomedical applications, the system could be used to clarify other types of nanoparticle probes, including tagged particles or molecules. Their research appears in the Proceedings of the National Academy of Sciences of the United States of America (PNAS).
“As more powerful imaging technologies are developed, scientists face a real challenge to quantitatively analyze and interpret these new mountains of data,” said Wang. “This paper is only a start, but I expect that innovative computing and data processing will be increasingly used to reveal detailed and quantitative features not currently available to biomedical researchers.”
“This work is pointing to a new era in light microscopy in which single-molecule detection is achieved at nanometer resolution,” added Nie. “This is also an example of interdisciplinary research in which advanced computing meets nanotechnology. I envision major applications not only for single-molecule imaging but also for ultrasensitive medical diagnostics.”
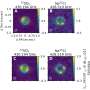
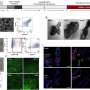
Because scientists frequently use several different colors of nanoprobes to color-code genes and proteins, a blended color dot is a common challenge when analyzing images. For every few green or red dots in an image, there could be a few yellow dots as well, indicating that at least two dots are clustering to create the appearance of a new color. Although less than precise nanoprobe images yield valuable information, the Georgia Tech and Emory research team knew that better technology was needed to pinpoint the exact distance in nanometers between probes to reveal important information about the size and binding geometry of targeted molecules.
“We had no way of knowing for sure if we were looking at one molecule or two or three molecules very near one another,” said Wang. “The fuzzy dot images were not precise enough on the nanometer level to truly tell us how these markers reflect DNA, but this system allows us to collect quantitative data and prove—not hypothesize—how genes are behaving.”
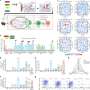
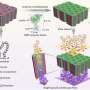
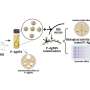
Instead of starting from scratch to create a system to isolate the clumped nanoprobe images, the Georgia Tech and Emory researchers pursued their stellar photometry idea by adapting DAOPHOT, a program written by Peter Stetson, Ph.D., at the Dominion Astrophysical Observatory, which was designed to handle crowded fields of stars. After adapting DAOPHOT, the research team used color-coded nanoparticles to beat the traditional diffraction limit by nearly two orders of magnitude, allowing routine superresolution imaging at 1-nanometer resolution. And by using DNA molecules, two color-coded nanoparticles are designed to recognize two binding sites on a single target. Then the particles are brought together within nanometer distances after target binding.
These distances are sorted out by highly efficient image processing technology, leading to the detection and identification of individual molecules based on the target’s geometric size. Compared with other single-molecule imaging methods, the Georgia Tech and Emory system allows for higher speed detection involving much larger sample volumes (microliter to milliliters).
This work, which was supported in part by the NCI’s Alliance for Nanotechnology in Cancer, is detailed in the paper “Nanometer-scale mapping and single-molecule detection with color-coded nanoparticle probes.” An investigator from Georgia State University also participated in this study. This paper is available through open access at the journal’s Web site (www.pnas.org/cgi/reprint/105/9/3298.pdf).
Source: National Cancer Institute