e-Science methods reveal new insights into antibiotic resistance
Large-scale computer simulations have pinpointed a tiny change in molecular structure that could account for drug resistance in Streptomices pneumoniae, the organism that causes childhood pneumonia and claims 3.5 million lives a year, mainly in developing countries. Such knowledge could be invaluable in designing new drugs that are effective against the drug resistant strain.
Experiments to find out how changes at the molecular level are causing this resistance are difficult and, so far, have not been done. Now, however, Peter Coveney and co-workers from UCL and Queen Mary, University of London have investigated the problem using computer modelling techniques. Their findings are amongst several outputs of the UK e-Science programme that are discussed in a special Theme Issue of Philosophical Transactions of the Royal Society A* which is published on 15 August.
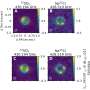
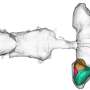
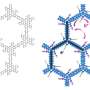
They took experimental data gathered from other organisms to build computer models of the sites where drug molecules interact with an organism’s protein molecules. They then ran simulations and visualised what happens when a drug molecule approaches each site for both normal and drug-resistant strains of S. pneumoniae.
The simulations and visualisations exploited highly scalable parallel code running on the UK’s national supercomputing facilities. “Without the use of e-Science methods, they would have taken months to perform and quite probably would never have been done. With these new methods, each simulation took just 12 hours,” says Professor Coveney. So far, life scientists have had limited access to and interest in such high performance computing resources; with Grid computing, these resources are becoming more readily accessible.
Professor Coveney and colleagues could see that a very small, but subtle, difference in structure between the normal and drug resistant strains was to blame for the drug resistance. In the normal strain, a drug molecule binds tightly to the site, but in the drug resistant strain it approaches and then drifts slowly away. If the results of the simulation are borne out by experiment, they could point the way to new drugs to combat disease.
*Large-scale molecular dynamics simulation of native and mutant dihydropteroate synthase-sulfanimide complexes suggests the basis of dihydropteroate synthase drug resistance by F.Giordanetto, P. W. Fowler, M Saqi and P. V. Coveney Philosophical Transactions of the Royal Society A 363 1833 (15 August 2005)
www.pubs.royalsoc.ac.uk/phil_t … scientificgrid.shtml