Scientists get first detailed look at Dicer
Scientists have gotten their first detailed look at the molecular structure of an enzyme that Nature has been using for eons to help silence unwanted genetic messages. A team of researchers with Lawrence Berkeley National Laboratory and the University of California at Berkeley used x-ray crystallography at Berkeley Lab's Advanced Light Source (ALS) to determine the crystal structure of Dicer, an enzyme that plays a critical role in the process known as RNA interference. The Dicer enzyme is able to snip a double-stranded form of RNA into segments that can attach themselves to genes and block their activity.
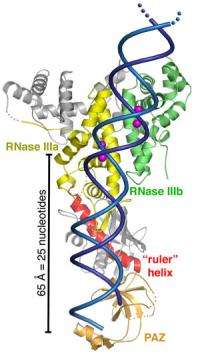
"With this crystal structure, we've learned that Dicer serves as a molecular ruler, with a clamp at one end and a cleaver at the other end a set distance away, that produces RNA fragments of an ideal size for gene-silencing," said Jennifer Doudna, a biochemist who led this study. Doudna, a leading authority on RNA molecular structures, holds joint appointments with Berkeley Lab's Physical Biosciences Division, UC Berkeley's Department of Molecular and Cell Biology and Department of Chemistry. She's also an investigator with the Howard Hughes Medical Institute (HHMI).
"Knowing the structure of Dicer sets the stage for understanding how Dicer enzymes are involved in other phases of the RNA interference pathway," Doudna said. "In human cells, the evidence points to Dicer being part of a larger molecular complex that directs the RNA interference process. The core structure of Dicer has been highly conserved by evolution and could serve as a guide in redesigning the RNA molecules that direct specific gene-silencing pathways."
RNA interference is an ancient gene-silencing process that plays a fundamental role in a number of important functions, including viral defense, chromatin remodeling, genome rearrangement, developmental timing, brain morphogenesis and stem cell maintenance. All of these RNA interference activities depend upon Dicer, so understanding this enzyme's molecular structure is a critical step.
The results of this research are reported in the January 13, 2006 edition of the journal Science in a paper entitled: Structural Basis for Double-Stranded RNA Processing by Dicer. Co-authoring the paper with Doudna were Ian MacRae, Kaihong Zhou, Fei Li, Adrian Repic, Angela Brooks, Zacheus Cande and Paul Adams.
RNA - ribonucleic acid – has long been known as a multipurpose biological workhorse, responsible for carrying DNA's genetic messages out from the nucleus of a living cell and using those messages to make specific proteins in a cell's cytoplasm. In 1998, however, scientists discovered that RNA can also block the synthesis of proteins from some of those genetic messages. This gene-silencing process is called RNA interference and it starts when a double-stranded segment of RNA encounters the enzyme Dicer. Double-stranded RNA (dsRNA) is formed from two single strands of RNA with complementary base sequences.
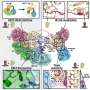
Dicer cleaves dsRNA into smaller fragments called short interfering RNAs (siRNAs) and microRNAs (miRNAs). Dicer then helps load these siRNA and miRNA fragments into a large multiprotein complex called RISC, for RNA-Induced Silencing Complex. RISC can seek out and capture messenger RNA (mRNA) molecules (the RNA that encodes the message of a gene) with a base sequence complementary to that of its siRNA or miRNA. This serves to either destroy the genetic message carried by the mRNA outright, or else block the subsequent synthesis of a protein.
Until now, it has not been known how Dicer is able to recognize dsRNA and cleave those molecules into products with lengths that are exactly what is needed to silence specific genes. Doudna and her co-authors were able to purify and crystallize a Dicer enzyme from Giardia intestinalis, a one-celled microscopic parasite that can infect the intestines of humans and animals. This Dicer enzyme in Giardia is identical to the core of a Dicer enzyme in higher eukaryotes, including humans, that cleaves dsRNA into lengths of about 25 bases.
In their Science paper, Doudna and her colleagues describe a front view of the structure as looking like an axe. On the handle end there is a domain that is known to bind to small RNA products, and on the blade end there is a domain that is able to cleave RNA. Between the clamp and the cleaver is a flat-surfaced region that carries a positive electrical charge. Doudna and her colleagues propose that this flat region binds to the negatively charged dsRNA like biological Velcro, enabling Dicer to measure out and snip specified lengths of siRNA.
"When you put the clamp, the flat area and the cleaver together, you get a pretty good idea as to how Dicer works," Doudna said. "We're now using this structural model to design experiments that might tell us what triggers Dicer into action."
Different forms of the Dicer enzyme are known to produce different lengths of siRNA. Having identified the flat-surfaced positively charged region in Dicer as the "ruler" portion of the enzyme, it may be possible to alter the length of a long connector helix within this domain to change the lengths of the resulting siRNA products.
"One size does not fit all for Dicer, it makes dsRNA products that range from 21 to 30 base pairs in length or longer. We would like to see what happens when you take a natural Dicer and change the length of its helix," Doudna said.
Determining Dicer's crystal structure was made possible through the unique crystallography capabilities of ALS Beamline 8.2.1 and 8.2.2, Doudna said. Funded through HHMI, beamlines 8.2.1 and 8.2.2 are powered by a superconducting bend magnet, an ideal source of x-rays for protein crystallography experiments. The "superbend" magnet is used to extract x-rays from a relativistic beam of electrons circulating through the ALS storage ring at energies up to two billion electron volts.
X-ray beams at the ALS are typically a hundred million times brighter than those from the best x-ray tubes. When a beam of x-rays is sent through a crystal, the atoms in the crystal cause the x-rays to scatter, creating a diffraction pattern. This diffraction pattern can be translated by computer into 3-D images of the crystal.
Source: Lawrence Berkeley National Laboratory