Bacteria which sense the Earth's magnetic field
Researchers uncover how a nanoscale 'compass' inside bacteria orients them to the Earth's magnetic field.
It is not only migratory birds that orient themselves to the magnetic field of the Earth. Also bacteria -- supposedly "simple" organisms -- have evolved to be able to take advantage of the magnetic field in their search for optimal living conditions. Such "magnetotactic"microorganisms use a miniature, cellular compass made of a chain of single nanomagnets, called magnetosomes.
The entire bacterium is oriented like a compass needle inside the magnetic field. Until now, it was not clear how the cells organise magnetosomes into a stable chain, against their physical tendency to collapse by magnetic attraction. But using modern molecular-genetic and imaging processes, researchers from the Max Planck Institue for Marine Microbiology in Bremen and Max Planck Institute of Biochemistry in Martinsried, Germany have identified the protein responsible for creating the magnetosome chain. The scientists showed that this protein aligns the magnetosomes along a cytoskeletal structure which was previously unknown. This points to evidence that genetics regulate the magnetosome chain exactly. The structure is one of the most complex that has ever been found in bacterial cells. It is comparable to organelles that, until now, scientists had only been familiar with in higher organisms. (Nature, Advanced Online Publication, November 20, 2005).
Magnetotactic bacteria are widespread in the mud of marine environments. In their cell interior, they form magnetosomes which are aligned into a chain. The bacteria use them to distinguish "up" from "down" in the Earth's magnetic field, and navigate themselves confidently through layers of water to efficiently find optimal growth conditions. The magnetosomes are made of tiny crystals of the magnetic iron mineral magnetite (Fe3O4) -- only about 50 nanometres in size.
To build magnetosomes, the cells do not only have to take up large amounts of iron from their surroundings and, from it, produce special iron oxide. The crystals also have to have a exactly defined number, form, and size, in order to be effective magnetic field sensors. To function optimally, the magnetosome crystals have to be strung into a straight chain inside the cell, thereby summing their magnetic moments. Only such a chain structure allows the magnetosomes to behave together like a compass needle which orients the bacteria inside the relatively weak magnetic field of the Earth. But until now, scientists did not know how magnetosome chains are formed.
The research group of Dr Dirk Schüler at the Max Planck Institute for Marine Microbiology has been investigating magnetosome formation in the magnetotactic bacterium Magnetospirillum gryphiswaldense, which the scientists found in the mud of a creek at Greifswald, in the northeast corner of Germany. Recently, the researchers were able to identify the part of the DNA that seemed to carry the entire genetic information required for formation and organisation of magnetosome particles. In this genomic fragment, known as a "magnetosome island", there are at least 25-30 different magnetosome genes, whose exact role had not been known in detail.
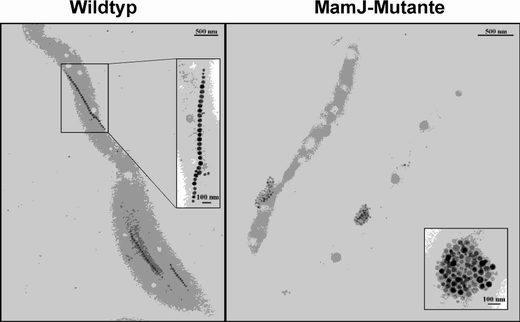
Magnetotactic bacteria compared. Left: genetically unaltered cells ("wild type") of Magnetospirillum gryphiswaldense contain as many as one-hundred magnetite crystals. They are nanosized permanent magnets, which are arranged into a nearly perfect chain inside the cell. The magnetosome chain orients the bacteria similarly to the way a compass needle in the Earth's magnetic field works. Right: a particular gene was removed from a mutant, resulting in the loss of the MamJ protein. The magnetosomes group together into irregularly ordered clusters. Here, because the magnetic moments of the individual magnetite crystals partially cancel each other out, the bacteria without MamJ can orient themselves only weakly in the magnetic field. Image: Max Planck Institute for Marine Microbiology
The researchers more closely investigated the magnetosome island and its function. They came across a gene, one of whose products (among other magnetosome proteins) is a component of the membrane that encloses every indivudual magnetite crystal. This protein, called MamJ, exhibits an unusually high portion of amino acids, aligned in repeats. MamJ has a remote similarity to proteins which control crystallisation processes in other biominerals, like bones, teeth, otoliths, and mussel shells. Thus, the scientists initially suspected MamJ is responsible for the development of magnetite crystals.
Although it is difficult to cultivate and manipulate Magnetospirillum gryphiswaldense in the laboratory, as part of his doctoral work Andrй Scheffel succeeded in removing the relevant gene from the genome. In this way, mutant bacteria were created which were lacking the MamJ protein. The mutants, surprisingly, still developed magnetosome crystals, which resembled the wild type in shape, size, and number. But the sensitivity of the magnetic field sensor function was disrupted; the cells could only orient themselves weakly in the magnetic field. Inspection by an electron microscope showed that the mutants' magnetosome crystals did not build a perfectly organised linear chain -- as the wild type's did -- but rather clumped together into irregularly arranged clusters. (see image)
The scientists genetically marked the MamJ protein with the fluorescent reporter protein GFP ("green fluorescent protein") and could thus track the fusion protein in the living bacteria cells. As they suspected, the protein was associated with the magnetosome chain. But microscopy made it apparent that MamJ arranges itself along a filamentous structure which trails like a string through the entire cell. In looking for this structure, the researchers used a new electron microscopic method which was developed in the Department of Molecular Structural Biology at the Max Planck Institute of Biochemistry, and which has already helped shed light on many cellular structures and functions.
Cyroelectron tomography makes it possible to analyse structures in detail inside an intact, shock-frozen cell (minus 196 degrees celsius), and to display them in three dimensions, at a resolution of just a few nanometres. In Dr Jürgen Plitzko's research group in Martinsried, Manuela Gruska's doctoral work involved investigating magnetic bacteria cells using this technology, and comparing the wild type with the MamJ mutants. She was able to make visible not only the magnetite crystal, but also the surrounding membrane vesicle, in a resolution never achieved before.
Amazingly, the scientists saw a previously unknown filamentous structure along the magnetosome chain of the wild type cells. It resembled a similar structure which structural biologists from Martinsried had already imaged in three dimensions in other cells. This shed light on the crux of the problem with the magnetosome chain: although in the wild type, the magnetosomes lay like pearls on a string along the filament, in the cells missing MamJ, the empty magnetosome vesicles scattered themselves about. It also explains why, as soon as they grow to a particular size, the magnetic crystals clump together in mutant bacteria.
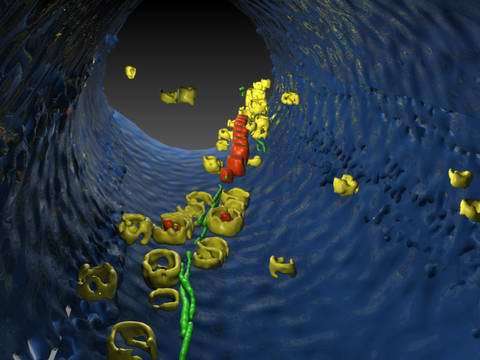
Image: Cryoelectron tomography of a magnetotactic bacterium: a three-dimensional reconstruction of the interior of a Magnetospirillum gryphiswaldense cell. The cell membrane is blue, the magnetosome crystal red, and the surrounding vesicle yellow. The image makes it clear that both the membrane vesicle and the "mature" magnetosomes are strung like pearls on a chain along a filamentous structure (green), which is similar to a cytoskeleton. Image: Max Planck Institute for Biochemistry
The scientists speculate that the MamJ protein develops, on one hand, on the surface of the magnetosome, and on the other hand, on the newly-discovered filament. It thus enables a close connection between the magnetosome vesicle and structure. This is further evidence of the diverse functions that cytoskeletal structures in bacteria seem to have, and which, for a long time, have only been known in eukaryotes -- i.e., organisms whose cells have nuclei.
The fact that the chain structure of bacterial nanomagnets is exactly regulated by genetics could also have implications for our understanding of the way higher organisms orient themselves to the magnetic field. We have known for some years that certain animals -- like migratory salmon or homing pigeons -- have magnetite crystal chains in particular tissue. These chains are astoundingly similar to those in bacteria, and possibly develop through a related mechanism.
Source: Max-Planck-Gesellschaft