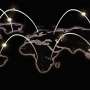
Twisted Molecule: Large and Folded Like Protein -- But Completely Synthetic
The physiological functions of proteins depend on their folding into a particular spatial structure (tertiary structure): enzymes and their substrates must fit together like the proverbial lock and key. It has recently been discovered that not only large biomolecules are capable of stable, defined folding; synthetic molecules can do it too. Called foldamers, these molecules can even imitate the biological functions of the proteins they are modeled after.
However, until recently their size and complexity was strictly limited. French researchers have now produced an intricately folded molecule exclusively from manmade components. The dimensions of this foldamer correspond to those of the tertiary structures of smaller proteins.
The team led by Ivan Huc did not want to base the design of their foldamer on the structure of proteins, because the synthesis of large chains from small individual building blocks is difficult. The alternative was to use branched structures. They did adopt one important structural element from proteins: the helix. The researchers hooked eight quinoline units (nitrogen-containing aromatic six-membered rings with a shared edge) together into a chain. This type of octamer twists itself into a helix.
The researchers then bridged two such octamers together with a special branching link. This linker inserts so well into the two octamers that a continuous, stable helix is formed. The branching linker can then be used to hook two such helical structures together side by side. Once linked, the two helices do not lie in parallel, but rather at right angles to each other.
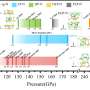
Helices can be twisted to the left or the right. In peptides, the direction of the helix is uniquely defined by the spatial structure of the individual building blocks. In the synthesis of the quadruple-octamers, however, an equal number of right- and left-handed helices are formed. The preferences demonstrated by the helices on pairing are determined by the solvent: In aromatic solvents, pairing of two helices with the same direction of twist is clearly preferred (70 %), while in chlorinated hydrocarbons up to 93 % of the pairs are formed from helices with opposite directions of twist. When the solvent is changed, the helices change their directionality to match these preferences.
“This proves both helices are involved in strong interactions with each other, just like a folded protein,” says Huc. “Our abiotic foldamer is the first of its kind and shows that it is possible to synthesize folded molecules that imitate the size and structural complexity of the tertiary structure of proteins, while consisting entirely of manmade building blocks.” The goal is to produce artificial structures with defined binding sites and uniquely positioned catalytic groups for controlled reactions with specific substrates.
Citation: Ivan Huc, Proteomorphous Objects from Abiotic Backbones, Angewandte Chemie International Edition, doi: 10.1002/anie.200603390
Source: Angewandte Chemie