The CNT-DNA wrap: A hefty hybrid for carbon nanotubes
Since their discovery in 1991, carbon nanotubes (CNTs) have captured the public imagination. These scrolls of graphite are much too tiny to be seen but they are stronger than diamonds. Formed from organic material, they can be shaped in a variety of ways and can act as metals or as semiconductors. They offer great potential in nanoelectronics, medicine, sensing and lasers, and as strengthening elements in composite materials.
Several obstacles must be overcome, however, before CNTs live up to their expectations. Chief among these is the tendency of CNTs to clump together like strands of angel-hair pasta. Other challenges include a better understanding of CNT structures, and more effective ways of processing the tubes, sorting them, placing them on substrates, and engineering their properties.
Lehigh University, in collaboration with DuPont and MIT, recently received a four-year, $1.25-million grant from the National Science Foundation to solve these problems by developing and studying new methods of manipulating CNTs in solution.
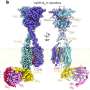
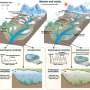
The Lehigh researchers will work with MIT, Cornell and DuPont through NSF's Nanoscale Interdisciplinary Research Team (NIRT) program and its Grant Opportunities for Academic Liaison with Industry (GOALI) initiative. Much of the team's focus will be on the use of single-walled CNTs wrapped with single-stranded DNA, a process that forms a helix around the nanotubes. The DNA-CNT hybrid has proven effective in CNT dispersion and researchers hope it will also aid in sorting and placing the tubes.
Several years ago, a DuPont-led research team found that DNA strands could be used to separate CNTs according to their electronic characteristics. The discovery was reported in an article in Science and cited later by Forbes magazine as one of the top five nanotechnology breakthroughs of 2003.
Principal investigators on the NIRT team include DuPont scientist Ming Zheng, Anand Jagota, formerly of DuPont and now the director of Lehigh's bioengineering and life sciences program, Slava Rotkin of Lehigh's physics department, Christopher Kiely of the Center for Advanced Materials and Nanotechnology at Lehigh, and Yet-Ming Chiang of the materials science and engineering department at MIT.
The two main goals of the NIRT team are to place CNTs on a substrate in specific locations and with specific densities and orientations, and to sort a heterogeneous sample of CNTs into constituent types.
To accomplish this, says Jagota, the NIRT team will seek to predict the structure of the DNA-CNT hybrid, given the DNA sequence and the CNT type, and to design experiments to control the placement and separation of the CNTs.
To gain greater control over the placing of CNTs on substrates, the researchers will apply a recently discovered technique called quasi-2D liquid crystal formation at a liquid-solid interface.
"If we can do what we're hoping to do," says Jagota, who is also a professor of chemical engineering at Lehigh, "we will have achieved a major advancement in CNT research."
Characterization for the NIRT project will be supervised by Kiely, professor of materials science and engineering and director of the Lehigh CAMN's Nanocharacterization Laboratory. The theoretical work will be overseen by Rotkin, assistant professor of physics at Lehigh. Both are co-PIs in the project.
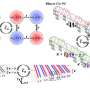
Rotkin is seeking to determine whether and to what degree the nanotube structure, specifically its bandgap structure, is altered when the CNT is wrapped by the DNA strand. He is using quantum-field analytical and numerical quantum-mechanical calculations to examine different types of CNTs with different types of DNA wraps.
"The answer to the question – does a CNT wrapped with DNA stay the same or undergo a change in properties? – depends on the symmetry and geometry of the wrap," says Rotkin.
"In some cases, the original CNT is metallic and has no bandgap. With the addition of the DNA strand, the CNT may or may not acquire a bandgap. The resulting hybrid properties and bandgap structure depend, first of all, on the original CNT properties and, second, on how the DNA is wrapped."
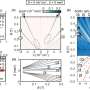
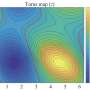
Rotkin and his students have succeeded in plotting bandgap structure, and mapping the areas of varying DNA-induced electric charges, which show up in repeating patterns.
"We more or less understand the rules of nature in regards to whether or not the bandgap structure [of a DNA-wrapped CNT] changes," he says. "But there are a huge number of nanotube types, and a huge number of ways of wrapping a CNT."
Source: Lehigh University