Gene scan helps identify cause of inherited blindness
(PhysOrg.com) -- Scientists at Washington University School of Medicine in St. Louis have scanned the entire genome of mice for genes that help build photoreceptors, the light-sensing cells of the eye.
The results have already helped researchers identify the gene that causes a form of retinitis pigmentosa, a type of inherited blindness in humans.
"Quite a few of the more than 160 genes linked to blindness are active in photoreceptor cells," says Joseph Corbo, MD, PhD, assistant professor of pathology and immunology. "There are many of these disease genes still left to be discovered, and when we find patients whose blindness can't be explained by genes we already know about, this new dataset can serve as a pointer to the best places to look."
Results of the gene scan appear this month in Genome Research; the identification of the retinitis pigmentosa gene is detailed in a separate paper published last month in The American Journal of Human Genetics.
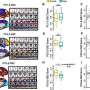
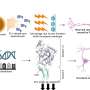
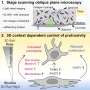
Corbo's lab and collaborator Thomas Langmann, PhD, of the Institute of Human Genetics in Regensburg, Germany, conducted the scan by looking for regions on mouse DNA that can bind to CRX, a protein that turns genes on or off in photoreceptors. They found CRX binding sites near many of the genes already known to be involved in photoreceptor development as well as hundreds of additional genes.
According to Corbo, the results indicate that CRX is a more important driver of photoreceptor development than researchers realized.
"We knew it was a key regulator, but we didn't realize the extent of its influence," he says. "It seems to have binding sites around almost every photoreceptor gene."
In a second study, researchers applied the scan results to genetic data from a family with an unexplained form of retinitis pigmentosa. Previous studies had determined that the problem in this family lies in a region of DNA containing 134 genes. Researchers used CRX binding to pinpoint a gene called FAM161A as the cause of blindness in this family.
"The CRX binding results allowed us to rapidly prioritize which of the 134 genes in this region were likely to be causing the disease." Corbo explains. "I think this is going to be widely applicable in the hunt for other blindness genes, which is important because finding the causative gene is a necessary step toward developing effective therapies for individual patients."
More information: -- Corbo JC, Lawrence KA, Karlstetter M, Myers CA, Abdelaziz M, Dirkes W, Weigelt K, Seifert M, Benes V, Fritsche LG, Weber BHF, Langmann T. CRX ChIP-seq reveals the cis-regulatory architecture of mouse photoreceptors. Genome Research, online Aug. 6, 2010.
-- Langmann T, Di Gioia SA, Rau I, Stöhr H, Maksimovic NS, Corbo JC, Renner AB, Zrenner E, Kumaramanickavel G, Karlstetter M, Arsenijevic Y, Weber BHF, Gal A, Rivolta C. Nonsense mutations in FAM161A cause RP28-associated recessive retinitis pigmentosa. The American Journal of Human Genetics (2010), doi:10.1016/j.ajhg.2010.07.018