Quantum dots spotlight DNA-repair proteins in motion
Repair proteins appear to efficiently scan the genome for errors by jumping like fleas between DNA molecules, sliding along the strands, and perhaps pausing at suspicious spots, say researchers at the University of Pittsburgh, the University of Essex and the University of Vermont who tagged the proteins with quantum dots to watch the action unfold. The findings are available today in Molecular Cell.
Everyone is constantly bombarded with environmental toxins that inflict small errors in the DNA code, so a rapid repair system is essential to maintain the integrity of the sequences for proper cell function, explained senior author Bennett Van Houten, Ph.D., Richard M. Cyert Professor of Molecular Oncology and leader, molecular and cellular cancer biology program, University of Pittsburgh Cancer Institute (UPCI), and professor, Department of Pharmacology and Chemical Biology, University of Pittsburgh School of Medicine.
"How this system works is an important unanswered question in this field," he said. "It has to be able to identify very small mistakes in a 3-dimensional morass of gene strands. It's akin to spotting potholes on every street all over the country and getting them fixed before the next rush hour."
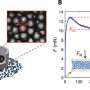
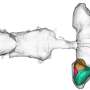
The researchers sought to unravel the mystery by tagging two repair proteins, called UvrA and UvrB, with quantum dots, which are semi-conductor nanocrystals that light up in different colors. They also stretched the usually clumped DNA into multiple "tightropes" to see the process more clearly.
They watched while UvrA proteins randomly jumped from one DNA molecule to the next, holding on to one spot for about seven seconds before hopping to another site. But when UvrA formed a complex with two UvrB molecules (UvrAB), a new and more efficient search technique emerged: the complex slid along the DNA tightrope for as long as 40 seconds before detaching itself and jumping to another molecule.
"If an E.coli bacterium had only one UvrAB complex, 13 hours would elapse before the entire genome was scanned for errors," said lead researcher Neil M. Kad, Ph.D., Department of Biological Sciences, University of Essex, United Kingdom. "About 40 complexes, comparable to the estimates of what occurs naturally, would be needed to scan it within the bacterium's 20-minute doubling time."
In addition to random jumping and sliding, the researchers also observed what they called "paused motion," in which UvrAB's motion seemed slower and purposeful.
"About one-third of the motile molecules in our study behaved this way," said co-author David M. Warshaw, Ph.D., professor and chair, Department of Molecular Physiology and Biophysics, University of Vermont. "Paused motion could represent UvrAB complexes checking for structural abnormalities associated with DNA damage."
The researchers now are exploring the possibility that the complexes sample the shape or chemical configuration of DNA by interacting with it; an error could alter the local DNA structure, changing its handshake with the repair proteins and perhaps triggering a corrective response.
Provided by University of Pittsburgh