Growth factor gene shown to be a key to cleft palate
Cleft palate has been linked to dozens of genes. During their investigation of one of these genes, researchers at Washington University School of Medicine in St. Louis were surprised to find that cleft palate occurs both when the gene is more active and when it is less active than normal.
They say the finding suggests this gene and processes closely associated with it are central to palate development and could become important targets for investigators seeking nonsurgical treatments to prevent cleft palate before birth. Their report will appear in an upcoming issue of the Proceedings of the National Academy of Sciences.
"Palate formation in the embryo is a complex process, and many things can go wrong," says senior author David M. Ornitz, M.D., Ph.D., Alumni Endowed Professor of Developmental Biology. "A cleft palate is often diagnosed late in pregnancy and treated surgically after birth. But if we understood the genetic causes of this common birth defect, we might be able to diagnose it much earlier. That would potentially allow intervention with prenatal surgery or with drugs or other agents designed to counteract the genetic abnormalities."
Clefts of the lip and palate affect about one in 700 newborns worldwide. Children with cleft lip and palate can have difficulty feeding as infants and can have speech, dental and hearing problems as they grow older. Depending on severity, surgical repair can require several operations over many years, and the estimated average lifetime cost of treatment in the United States is about $100,000 per patient.
"We believe the more information we have on the causes of cleft palate, the better hope we have for possibly preventing and more effectively treating the condition," says lead author Alison K. Snyder-Warwick, M.D., a plastic surgery resident at Barnes-Jewish Hospital.
Although some cases of cleft lip and palate are linked to environmental factors such as maternal smoking, viral infections or certain medications, genetic variations play a significant role in many cases. The Washington University researchers studied the fibroblast growth factor receptor 2 (FGFR2) gene, which earlier research has implicated in cleft palate.
They focused on mice with Crouzon syndrome, a developmental disorder caused by a mutation in FGFR2. The mutation activates the receptor and results in a syndrome that is characterized by abnormal development of the skull, face and mouth and is associated with an increased incidence of cleft palate.
In effect, the FGFR2 mutation prevents specific growth signals from being switched off. Normally, the signals would be turned on and off in a carefully orchestrated manner to ensure proper patterns of growth and development of embryonic tissues. However, the Crouzon syndrome mutation locks the receptor in a permanently on position.
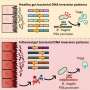
As mouse embryos with the mutation grew, cells destined to become the palate initially grew faster than normal cells, as anticipated. But just before palate formation, the growth of these cells lagged behind their normal pace of proliferation. That was unexpected because the signals created by mutant FGFR2 should logically have maintained an increased rate of cell proliferation in the palate, Ornitz indicates.
In a normal mouse embryo, groups of cells called the palatal shelf on either side of the mouth grow outward, elevate to meet in the middle and fuse to form the palate. But in the mutant mice embryos, the stunted growth of this tissue prevented the palatal shelves from properly elevating, meeting and fusing. In addition, the researchers detected a decrease in some key components of the supporting matrix between cells of the palate.
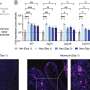
Another of the study's coauthors, Kai Yu, Ph.D., a scientist in the Ornitz lab, created genetically engineered mice in which FGF receptors were inactivated in tissue that gives rise to the palate. These mice also developed cleft palate.
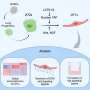
In palate cells grown in the lab, the researchers looked at the FGF cell-signaling network, in which FGFR2 participates. They compared the effect of increased activity of FGF signaling with decreased activity of the same network, and interestingly, both led to cleft palate.
"We found that overactivation of an important signaling pathway resulted in loss of function," Snyder-Warwick says. "Our results suggest a different way of thinking about mutations in the FGF signaling network. This study clearly showed that this FGF signaling pathway is a critical regulator of palate development."
These findings strengthen the evidence that the developmental processes in which FGFR2 is involved could be targeted with drugs in order to ensure normal palate growth.
"It's possible that someday doctors might be able to administer drugs that would either slightly activate or slightly inhibit FGFR2 function," Ornitz says. "That might be enough to tip the balance from a cleft palate to a normal palate during embryonic development."
More information: Snyder-Warwick AK, Perlyn CA, Pan J, Yu K, Zhang L, Ornitz DM. A gain-of-function FGFR2 Crouzon mutation: Evidence of loss-of-function activity in the etiology of cleft palate. Proceedings of the National Academy of Sciences (advance online publication).