From fruit fly wings to heart failure -- why Not(ch)?
Almost a century after it was discovered in fruit flies with notches in their wings, the Notch signalling pathway may come to play an important role in the recovery from heart attacks. In a study published today in Circulation Research, scientists at the European Molecular Biology Laboratory (EMBL) in Monterotondo, Italy, are the first to prove that this signalling pathway targets heart muscle cells and thus reveal its crucial role in heart development and repair.
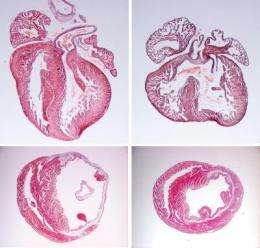
The Notch pathway is a molecular mechanism through which cells communicate with each other. Scientists in Nadia Rosenthal's group at EMBL used sophisticated genetic mouse models to uncover critical roles for this pathway in heart muscle cells. When they inactivated Notch specifically in the heart muscle precursor cells of early mouse embryos, the scientists discovered that the mice developed heart defects.
Curiously, increasing Notch signalling in the heart muscle cells of older embryos had the same detrimental effect, uncovering different requirements for Notch as development proceeds.
"The cardiac malformations we observed are characteristic of Alagille syndrome, a human congenital disorder," said first author Paschalis Kratsios. "Therefore, our findings could help to explain the cardiac symptoms associated with Alagille syndrome and related forms of congenital heart disease."
Intriguingly, the scientists were able to improve the cardiac function and survival rate of adult mice that had suffered heart attacks by re-activating Notch, suggesting new therapeutic approaches to help the heart recover from damage.
"Overall, these results highlight the importance of timing and context in biological communication mechanisms" Nadia Rosenthal concludes: "Our findings also lend support to the notion that, in certain situations, redeployment of embryonic signalling pathways could prove beneficial for tissue regeneration in the adult."
Source: European Molecular Biology Laboratory (news : web)