New insights into cardiac aging
Investigators at Burnham Institute for Medical Research (Burnham) have found that the conserved protein d4eBP modulates cardiac aging in Drosophila (fruit flies). The team also found that d4eBP, which binds to the protein dEif4e, protects heart function against aging. This research enhances our understanding of the TOR and FoxO signaling pathways and provides a more specific target for further research into cardiac aging. Since the TOR and FoxO genes are conserved between Drosophila and humans, this work may lead to new, tissue-specific methods to protect the heart. The paper was published in the journal Aging Cell.
Much research has shown that altering the expression of specific genes can extend the lifespan of various organisms. Overexpression of dFoxO and reduced expression of dTOR both work to extend Drosophila lifespan. However, researchers needed to investigate the mechanisms behind these pathways, as well as how these signaling pathways influence aging in specific tissues, in this case the heart.
"The relationships between these genes are very complex," said Rolf Bodmer, Ph.D., who directs Burnham's Development and Aging Program. "We wanted to analyze how two opposing genes function and control their downstream effectors, and we wanted to understand how these aging factors apply to a specific organ."
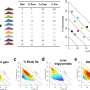
The Bodmer laboratory, in collaboration with the laboratory of Sean Oldham, Ph.D., an expert in TOR signaling, altered the expression levels of dTOR pathway components in heart tissue and tested the hearts' stress response. Increased dTOR activation resulted in higher failure rates, while reductions in dTOR activity promoted more youthful hearts. Noting that upregulated dFoxO and downregulated dTOR lead to similar consequences, the laboratory looked for downstream factors that were influenced by both pathways. One possibility was d4eBP, which reduces messenger RNA translation by binding to dEif4e. The team found that increased d4eBP levels produced the same healthier hearts as decreased dTOR activity, while increased dEif4e levels resulted in higher failure rates.
The team also showed that when dTOR and its antagonistic effecter d4eBP were co-expressed, the hearts did not differ significantly from when d4eBP was expressed by itself, indicating that there is a straight signaling path from dTOR to d4eBP/dEif4e. These new findings also introduce the interesting biological concept that changes in (TOR-dependent) mRNA translation factors (d4eBP and dEif4e) influence the age-dependent functional performance of the heart.
Source: Burnham Institute (news : web)