Genes key to staph disease severity, drug resistance found hitchhiking together
Scientists studying Staphylococcus bacteria, including methicillin-resistant S. aureus (MRSA), have discovered a potent staph toxin responsible for disease severity. They also found the gene for the toxin traveling with a genetic component of Staphylococcus that controls resistance to antibiotics. The study, now online in PLoS Pathogens, shows for the first time that genetic factors that affect Staphylococcus virulence and drug resistance can be transferred from one strain to another in one exchange event.
One of the ways Staphylococcus bacteria become drug-resistant is through horizontal gene transfer, whereby resistance genes move from one bacterium to another. Staph bacteria also can exchange virulence genes using the same mechanism, but this was previously assumed to occur separately from the transfer of antibiotic resistance.
Scientists from the National Institute of Allergy and Infectious Diseases (NIAID), a component of the National Institutes of Health, led the study. They collaborated with researchers at the University of Tubingen in Germany and the University of Medicine and Dentistry of New Jersey.
"The discovery that bundled genes determine virulence and antimicrobial resistance suggests a new research focus for scientists trying to better prevent and treat serious staph infections," says Anthony S. Fauci, M.D., NIAID director.
The research involved more than 100 strains of S. aureus and S. epidermidis, both bacteria found on the skin of most people. In recent decades, these bacteria have become increasingly virulent, often causing severe disease that can be resistant to traditional antibiotics such as methicillin.
The studies were directed by NIAID senior investigator Michael Otto, Ph.D. In 2007, he and his colleagues found that staphylococci secrete toxins of the phenol-soluble modulin (PSM) family that are primarily responsible for attracting and killing human white blood cells called neutrophils. This process is critical for the ability of S. aureus—including community-acquired MRSA—to cause disease.
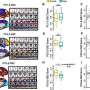
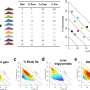
While screening S. aureus and S. epidermidis strains, Dr. Otto's group noticed that some strains produced one additional, previously unknown, PSM toxin. The researchers hypothesized that the toxin was somehow connected to drug resistance. This idea surfaced because the toxin appeared in 10 percent of all MRSA strains and 68 percent of all methicillin-resistant S. epidermidis strains analyzed—whereas the researchers did not find it in strains of S. aureus or S. epidermidis that are sensitive to methicillin.
The research group confirmed its theory by identifying the specific location that encodes the toxin, which was in gene clusters that control drug resistance, known as SCCmec. The group named the new toxin PSM-mec.
"This work represents a previously unknown example of a toxin hitchhiking on staphylococcal mobile genetic elements that are primarily in charge of transferring antibiotic resistance," says Dr. Otto. He adds that the finding "should alert the research community that aggressive, drug-resistant staph can evolve more quickly than we assumed."
The research group is continuing its study of PSM-mec in S. epidermidis, where the toxin is more prevalent. Ultimately, being able to neutralize PSM-mec and other toxins that attack human defenses could lead to new treatments for S. aureus and S. epidermidis disease.
More information:
S Queck et al. Mobile genetic element-encoded cytolysin connects virulence to methicillin resistance in MRSA. PLoS Pathogens DOI: 10.1371/journal.ppat.1000533 (2009).
R Wang et al. Identification of novel cytolytic peptides as keyDOI: 10.1038/nm1656ants of community-associated MRSA. Nature Medicine DOI: 10.1038/nm1656 (2007).
Source: NIH/National Institute of Allergy and Infectious Diseases (news : web)