Covering the bases: Quantum effect may hold promise for low-cost DNA sequencing, sensor applications
A ghostly property of matter, called quantum tunneling, may aid the quest for accurate, low-cost genomic sequencing, according to a new paper in Nature Nanotechnology Letters by Stuart Lindsay and his collaborators at the Biodesign Institute of Arizona State University. Tunneling implies that a particle, say an electron, can cross a barrier, when, according to classical physics, it does not have enough energy to do so.
Unraveling the DNA sequences of the human genome a decade ago was a remarkable achievement. Today, the task of sequencing some 3 billion chemical base pairs of the genome—enough information to fill a 20-volume encyclopedia—remains a daunting challenge, thus far accomplished largely through brute force means. Such methods are typically slow and extravagantly expensive, (though costs have dropped considerably from the initial sequencing of the human genome, which took 11 years at a cost of $1 billion.)
Bringing the power of DNA sequencing to every individual will require new, affordable technologies to help mine the wealth of information DNA can provide concerning morphology, hereditary traits and predisposition to disease.
Various techniques for sequencing DNA have been used to determine the identities of the four nucleotide bases—adenine, thymine, cytosine and guanine—which make up the ladder rungs of the DNA's double helical structure. Most of these require snipping DNA into hundreds of thousands of short fragments, unzipping the helix and reading a few hundred to a few thousand bases at a time. Finally, all of the information from the DNA pieces is reassembled into a picture of the complete genome, with the help of massive computing power.
ASU Regents' Professor and Carson Presidential Chair of Physics and Chemistry, Stuart Lindsay, who also directs the Biodesign Institute's Center for Single Molecule Biophysics, summarizes one of the chief physical obstacles to more efficient identification of DNA base pairs through techniques like optical microscopy: "The difficulty is that any physical readout that you can think of placing on a device is sensitive on a length scale that is longer than the separation between bases."
Lindsay believes there may be a radical solution.
The rules of attraction
Dr. Lindsay's technique for observing DNA sequences relies on devices known as scanning tunneling- (STM) and atomic force- (ATM) microscopes. He exploits these sensitive instruments to identify complementary DNA base pairs, evaluating the hydrogen bonds formed between them. Base pairing rules for DNA dictate that the hydrogen bonds work to join up appropriate nucleotide pairs like jigsaw pieces—adenine with thymine and cytosine with guanine.
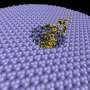
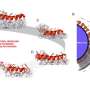
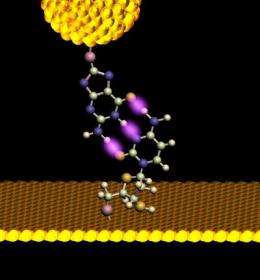
The scanning tunneling microscope used in the present study features a delicate electrode tip held very close to the DNA sample. When this tip is fitted with a particular nucleotide and brought in contact with its complementary mate—embedded in the substrate, the hydrogen bonds stick the bases together and they attach, like tiny magnets (see fig. 1). As Lindsay describes the method, " you have sensing chemicals attached to one electrode and the target you want to sense attached to another one. When the junction spontaneously self-assembles, you get a signal. It's a new way of doing recognition at the atomic scale."
Crucial to the new technique is the fact that the strength of the glue fastening complementary bases differs for A-T and C-G pairs. While two hydrogen bonds hold A-T bases together, C-G pairs use three hydrogen bonds. For this reason, it's physically harder to break C-G bonds. By measuring the current drop in the electrical circuit formed between the microscope probe and the substrate as the hydrogen bonds are gently pulled apart, a positive identification of the base being read can be made. The new method, as Lindsay explains, combines chemical recognition—the hydrogen bonded assembly at the tunnel junction— with the flow of electron tunneling current as the tunneling junction is completed.
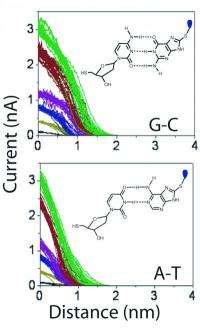
The study made a number of measurements using varying amounts of electrical current through the junction, as the microscope's electrode is moved away from the substrate and the hydrogen bonds uniting base pairs are slowly pulled apart. "What you can see straight away," Lindsay notes, "is that with the bases held together by 3 hydrogen bonds, the curves of falling current go on for a long distance. In those held together with two, they go on for less distance." (see fig. 2)
Bridging the divide
Electron tunneling is a peculiar property of matter acting over tiny distances at the atomic or subatomic scale. In a classical electric circuit, a gate is either open or closed, permitting or blocking the flow of current. But, as Lindsay explains, "when you start to get two electrodes so close to one another that they are within a few atomic diameters, then the electrons can actually leak from one electrode to the other, because in quantum mechanics, they're not confined." These electrons, which violate classical mechanics as they hop across the tiny junction, are said to "tunnel."
Now, Lindsay's research team has developed a method to identify different DNA base pairs, which could serve as the foundation for a new DNA sequencing technology. "The tunnel current is there as a readout of how long that molecular pair survived in the junction, " Lindsay says. "But it turns out that it's an incredibly nice way of identifying which molecular pair it was." Although quantum tunneling seems exotic, Lindsay points out that the routine leaking of electrons from one atom to another to form a chemical bond is a similar process.
If significant challenges to reading single molecules through such a technique can be overcome, the method holds the potential for inexpensive DNA sequencing, operating at the breakneck pace of thousands of base pairs per second. As professor Lindsay notes, "this combination of tunneling plus the chemistry is very powerful."
Source: Arizona State University (news : web)