First glimpse of a key DNA repair protein at work

Repairing breaks in the two strands of the DNA double helix is critical for avoiding cancer. In humans and other organisms, a molecular machine called the MRN complex is responsible for finding and signaling double-strand breaks (DSBs), then launching the error-free method of DNA repair called homologous recombination.
In an article in the online October 3, 2008 issue of the journal Cell, John Tainer of the Life Sciences Division at the U.S. Department of Energy's Lawrence Berkeley National Laboratory, leading a team of his colleagues from the Scripps Research Institute, reveals how the central component of the MRN complex performs its essential functions.
The core of the MRN complex is the protein Mre11 (M stands for Mre11, R for the Rad50 protein, and N for the Nbs1 protein). But without good models of how Mre11 interacts with DNA, based on evidence from high-resolution images, it was impossible to know how Mre11 recognizes the ends of the broken DNA and how – or even whether – it remodels DNA sequences. So just how Mre11 works has been a matter of dispute. Tainer and his colleague resolved these issues by going beyond images of the stand-alone structure of Mre11 to study Mre11 bound to DNA during the first steps of DSB repair.
"This breakthrough was possible because combined project efforts, funded by DOE to characterize microbial complexes and by the National Cancer Institute to examine complexes relevant to cancer, came together to define these DNA complexes," says Tainer. "Understanding how the body responds to DNA damage is fundamental for many potential cancer interventions and gene therapies. These results are especially exciting, as they open the door to the predetermined control of homologous recombination for cancer therapeutics and gene targeting."
The power of homologous recombination
Homologous recombination is important not only for error-free repair of DSBs but also for targeting genes to specific sites in the genome, a valuable technique in cell biology and a highly promising method for gene therapy and other emerging medical interventions.
Double-strand breaks occur naturally about ten times a day in every human cell; they are one of the most highly toxic and mutagenic kinds of DNA damage, implicated in cancer and many other diseases. Because cancer cells are more susceptible to damage than many normal cells when dividing, DSBs are created deliberately in most cancer therapies to kill dividing tumor cells. But the deliberate inactivation of DNA repair pathways may be an even more powerful method of killing cancer cells, because cancer cells often have defects in repair or signaling of DNA damage which would cripple them if left unrepaired.
Thus the MRN complex, a crucial component of the homologous-recombination method of DSB repair, is a key target for both new cancer therapeutics and for controlled gene targeting to treat inherited disorders in humans as well, plus having numerous other research applications.
What the Tainer team reports in Cell is that Mre11 not only finds the ends of broken DNA strands and links them together but also remodels the DNA in preparation for the work of other repair proteins, an essential function of Mre11 on which successful homologous recombination depends. Additional studies allowed the researchers to determine how mutations of the mre11 gene affect the protein's ability to do its job and in some cases to cause disease.
Prompt and accurate DNA repair is so essential to life that many of the molecular machines that perform DNA repair have changed little throughout the long course of evolution; today these machines show remarkable similarity in organisms as diverse as archaea, yeast, frogs, and human beings.
One consequence is that information about how Mre11 works in humans can be derived from organisms in which proteins and protein complexes are much more stable at room temperature than human proteins would be. For example, the extremophile Pyrococcus furiosus is an archaeal microorganism that grows happily at the temperature of boiling water; at room temperature, P. furiosus's proteins and their complexes are so stable they might as well be frozen.
Synapses and branching forks
Scott Williams and Gabriel Moncalian of the Scripps Institute's Department of Molecular Biology and the Skaggs Institute for Chemical Biology crystallized and determined x-ray crystal structures of key domains of P. furiosus Mre11 bound to DNA in two configurations, representing two kinds of DSB-repair interactions.
At beamline 12.3.1 at Berkeley Lab's Advanced Light Source, known as SIBYLS ("Structurally Integrated BiologY for Life Sciences"), the researchers performed both high-resolution protein crystallography and small-angle x-ray scattering (SAXS), which can image proteins and complexes in aqueous solution, closer to their natural state.
"Like breaking a piece of wood into two parts, the severed termini of the DNA in a double-strand break have ends of varying shape," says Williams. "The high-resolution snapshots we obtained of Mre11 binding to different DNA end-structures show how the Mre11 protein has evolved to hone in on and sense these variable mutagenic lesions."
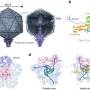
Of the two complexes that were crystallized, the "synaptic complex" bound the ends of a pair of DNA helices in orientations mimicking the kind of break that might result when a double strand of DNA is severed by radiation or chemical damage.
A different form of double-strand break is also common and occurs when a DNA helix is forking or "unzipping" during replication; if the process is interrupted, the fork may collapse. This "branched complex" of DNA bound with Mre11 involves different topography and interactions than the synaptic complex.
What the researchers found is that in both complexes two Mre11 proteins worked together as a dimer, forming a U-shaped structure. In the synaptic complex, the U-shaped dimer binds to two DNA double helices. The branching complex binds to the single helix that is unraveling at the point where forking has been interrupted.
In both cases binding occurs on the inner sides of the U, where both Mre11 subunits contribute to holding and manipulating each piece of DNA by means of "recognition loops" in the protein's folded string of amino acid residues. P. furiosus Mre11 has six recognition loops. Mre11 from eukaryotes – organisms including yeast, frogs, and humans, whose cells, unlike archaea and bacteria, have membrane-packaged nuclei – exhibit only five of these recognition loops but are otherwise similar.
How Mre11 works
The ends of the two DNA helices in the synaptic complex are held in close proximity while one part of each Mre11 protein, called the capping domain, rotates into place to adjust the position of the helices and prepare their ends. Another part of each subunit, called the nuclease domain, accesses individual nucleotides, stripping a strand from one of the helices to leave a so-called 3' tail, essential to the next steps in homologous recombination. The other strand, the 5' tail, is degraded in an Mre11 process known as a "resection," which also depends on nuclease activity.
The branching complex handles the different configuration of a collapsed fork in a different way. Where capping and nuclease domains meet, the twin Mre11s clamp onto the single helix where its division was halted. The 3' strand is held apart and prepared for continuing replication after repair.
Now combining sequencing and biochemical studies of Schizosaccharomyces pombe (fission yeast), by Paul Russell and his lab at Scripps, with models based on images of the P. furiosus Mre11-DNA complexes, the researchers learned how certain amino acid residues in the folded chains of the twin subunits interact with the DNA. With this knowledge they were able to explain how mutations of key amino acids in the sequence cause problems with DSB repair and can lead to disease, including rare disorders like Ataxia-telangiectasia-like disorder and Nijmegen breakage syndrome, which render the patient extremely vulnerable to ionizing radiation and genomic instability.
Mutations in two locations occupied by leucines, for example, inhibit self-interaction between the Mre11 proteins and the formation of the U-shaped dimer; as a result, Mre11's ability to bind to DNA is reduced and interactions with the other MRN proteins, Rad50 and Nbs1, can be affected.
On the other hand, mutations at two histidine sites do not affect the ability of Mre11 to form dimers or interact with Rad50, Nbs1, or other repair proteins, but do affect its ability to remodel DNA. One mutation, which affects the remodeling of DNA ends, causes fewer problems. The other histidine mutation impairs Mre11's ability to remodel both end and internal sequences and subsequently cripples repair. Because MRN initiates only one kind of DSB repair, however – homologous recombination – in some species a different kind of repair called nonhomologous end-joining can take up the slack.
The right tools for the job
DNA was bound to Mre11 dimers in the correct orientation and crystallized at the Scripps Research Institute in La Jolla, CA. Critical for the results was the analysis of the samples performed by the Scripps Institute's Williams at the SIBYLS beamline at Berkeley Lab's Advanced Light Source.
The SIBYLS beamline, 12.3.1, has unique capabilities to analyze the shape of samples in solution using small-angle x-ray scattering (SAXS), as well as details of samples in crystals using macromolecular crystallography (MX). Scott Classen of Berkeley Lab's Physical Biosciences Division (PBD) helped develop and is in charge of the MX facility, and Greg Hura, also of PBD, helped develop and is in charge of the SAXS facility. The Scripps Institute's Williams was sometimes able to work by remote control from La Jolla using the advanced automation at the SIBYLS end station, whose robots and software can load samples, track ongoing experiments in real time, and collect data.
"Some difficult-to-crystallize macromolecular complexes can result in large unit cells that are hard to work with. The MX endstation of SIBYLS is designed to address these challenges, for example with a large-area detector that can be positioned very far from the sample," says Classen. "While crystallography can give you nice pictures of static molecules, however, it isn't as informative when it comes to dynamic processes. That's where SAXS comes in; it's a perfect complement to MX."
Says Hura, "SAXS is a solution-imaging technique, where each experiment measures the sum of all conformations of the molecule. While the ultimate resolution of crystallography is higher, SAXS may be used to determine the flexibility of proteins as they perform their functions – and it can resolve the ambiguities of crystallography by showing what possible structures are the most likely."
Tainer, leader of SIBYLS, says "You get information from every protein crystal you analyze – if only what not to try crystallizing. Sometimes you can only crystallize a small part of a large protein or complex. It's the synergy with SAXS, which can show you how these pieces fit together or provide experimental evidence for the best among competing models, that makes this beamline combining both techniques so very valuable to the missions of both NCI and DOE."
Citation: "Mre11 dimers coordinate DNA end-bridging and nuclease processing in double-strand break repair," by R. Scott Williams, Gabriel Moncalian, Jessica S. Williams, Yoshiki Yamada, Oliver Limbo, David S. Shin, Lynda M. Groocock, Dana Cahill, Chiharu Hitomi1, Grant Guenther, Davide Moiani, James P. Carney, Paul Russell, and John A. Tainer, appears in the 3 October 2008 issue of Cell and is available online to subscribers at www.cell.com/content/article/a … PIIS0092867408010623 .
Source: Lawrence Berkeley National Laboratory