Plant Scientists Develop New Cell-Sorting Technique
A new cell-sorting technique developed by University of Arizona plant scientists has the potential to enhance our understanding of how cells of all types work – or, in the case of diseases such as cancer, how they fail to work.
Living organisms, including plants and animals, are made up of recognizable structures are called organs, that have specific functions – plant organs include leaves, roots, flowers and fruit, for example, while animals have kidneys, livers, hearts, brains and so on. Each of these organs in turn is made up of a complex mix of different cell types, each having different functions.
One of the challenges in understanding how organs function is the need to separate out these different cell types in order to analyze them. Other cell-sorting methods often rely on using a machine that recognizes the differences in the optical properties of the different cell types. However, it is difficult, and in some cases impossible, to separate the different cells within organs by this method.
UA plant sciences professor and BIO5 Institute member David Galbraith explains that instead of focusing on the cell as a whole, the technique his lab has developed focuses on cell nuclei. Other methods of teasing different cell types apart require that the cell remain intact – a challenge when working with plants that have rigid cell walls – and also when working with brain and other animal cells, which can be connected in intricate ways. "We thought it'd be a lot easier if we could take a razor and chop up the cells and then sort their nuclei," Galbraith said.
The nuclei of cells contain their genetic material – the "operating instructions" that regulate everything from how a saguaro stores water to how a human heart beats. For any given organism, the nucleus of every cell contains the same comprehensive set of instructions, but not every cell "reads" everything written there.
Heart cells don't carry out the instructions for digesting food, for instance; a tree's leaves don't carry out the instructions for drawing water out of the soil. Yet even a single leaf is made up of many cells with many subtly different functions.
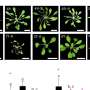
To develop their technique for separating such interconnected yet different cells, Galbraith and his graduate students focused on certain nutrient-transporting root cells in the small flowering plant Arabidopsis thaliana.
They modified a green fluorescent protein, or GFP, found in jellyfish by combining it with a protein that is found only in the nuclei of those root cells. They then arranged for this modified GFP to be produced only within specific cell types in the root, and as a result the GFP lit up only the nuclei within the cell types that the researchers were seeking.
They were then able to take advantage of GFP's visible fluorescence to allow the cell sorter to rapidly sort and purify the fluorescent nuclei of that cell type, while leaving the nuclei of other cells behind. Galbraith and his students further showed that the sorted nuclei provided the same information about the state of the “operating instructions” as that obtained from separated cells.
"Isolating nuclei is a simple alternative to isolating the whole cell," Galbraith said, "one that lets people analyze a lot of cell types they weren't previously able to analyze." In other words, scientists now have one more tool for understanding how cells work – in both plants and animals.
"This is an interdisciplinary kind of technology," Galbraith said. "The BIO5 Institute was an ideal place to develop this because it can now be applied not only to plants but also to animals and to biomedical problems."
While the individual research teams who make use of the new cell-sorting technique will decide what questions to tackle with it, Galbraith says this could potentially help us understand everything from how plants respond to environmental stresses and how to increase the yield of food crops, to how to treat diseases where the action of cells goes awry, as happens with cancer. "The more information we have, the more likely we are to be able to deal with problems," Galbraith said.
Source: University of Arizona