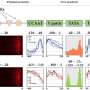
Flowering plants evolved very quickly into 5 groups

University of Florida and University of Texas at Austin scientists have shed light on what Charles Darwin called the “abominable mystery” of early plant evolution.
In two papers set to be published next week in the Proceedings of the National Academy of Sciences, the scientists report that the two largest groups of flowering plants are more closely related to each other than any of the other major lineages. These are the monocots, which include grasses and their relatives, and the eudicots, which include sunflowers and tomatoes.
Doug and Pam Soltis, a UF professor of botany and curator at UF’s Florida Museum of Natural History, respectively, also showed that a stunning diversification of flowering plants they are referring to as the “Big Bang” took place in the comparatively short period of less than 5 million years -- and resulted in all five major lineages of flowering plants that exist today.
“Flowering plants today comprise around 400,000 species,” said Pam Soltis. “So to think that the burst that give rise to almost all of these plants occurred in less than 5 million years is pretty amazing -- especially when you consider that flowering plants as a group have been around for at least 130 million years.”
The lead author of the UF paper is Michael Moore, a former postdoctoral associate in the Soltis lab and current faculty member at Oberlin College. Charles Bell, the fourth author, is another former Soltis postdoctoral associate, now at the University of New Orleans.
Robert Jansen, professor of integrative biology at The University of Texas at Austin, said the two papers set the stage for all future comparative studies of flowering plants.
“If you are interested in understanding the evolution of flowering plants, you can’t do that unless you understand their relationships,” he said.
Botanists predating Darwin have long recognized that flowering plants, which comprise at least 60 percent of all green plant species, diversified abruptly shortly after they appeared.
The details, and especially the cause of, this diversification – Darwin’s “abominable mystery” – has been a hot topic in botany ever since.
“One of the reasons why it’s been hard to understand evolutionary relationships among the major groups of flowering plants is because they diversified over such a short time frame,” Jansen said.
Seeking to distill the cloudy picture into a clear one, the UF and UT researchers analyzed DNA sequences from the completely sequenced genomes of the chloroplast. That organelle, responsible for plants’ ability to photosynthesize, is shared by all green plants.
Jansen and his UT Austin colleagues analyzed DNA sequences of 81 genes from the chloroplast genome of 64 plant species, while the UF researchers analyzed 61 genes from 45 species. The two groups also performed a combined analysis, which produced evolutionary trees that included all the major groups of flowering plants.
Assisting the effort at UF was a newly purchased rapid gene sequencing machine.
“UF was the first university to purchase this particular type of sequencing machine,” Doug Soltis said. “Where it would have taken months to sequence the chloroplast genome before, you can do it in a single day now.”
By laboriously arranging the sequences, the researchers slowly built a kind of family tree for plants – a diagram of relationships among plant lineages showing diversification over the eons. Based on known rates of genetic change double-checked against fossils of known ages, they established a time scale that revealed the dates of major branching events.
Based on the Soltises’ and their collaborators’ research in previous years, it was known that flowering plants split into three branches shortly after they appeared about 130 million years ago. That process was relatively gradual, at least compared with the rapid radiation that happened next. The details of that radiation have long been murky. The latest research clears the picture by showing that all plants fall into five major lineages that developed over the relatively short period of 5 million years, or possibly even less.
As for the diversification’s cause, it remains mysterious, Pam and Doug Soltis said.
It’s possible it was spurred by some major climatic event. It’s also possible that a new evolutionary trait – a water-conducting cell that transfers water up plant stems – proved so effective that it spurred massive plant species diversification. The cell is either not present, or is poorly developed, in the first three flowering plant lineages, Doug Soltis said. The earliest flowering plant lineages also did not have a completely fused ovary, which in later flowering plants may better protect the seeds, Pam Soltis said.
Source: University of Florida