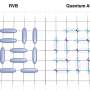
Prediction of RNA pseudoknots using heuristic modeling with mapping and sequential folding
An algorithm utilizing structure mapping and thermodynamics is introduced for RNA pseudoknot prediction. The method finds the minimum free energy in the context of the biological folding direction (5’ to 3’) of RNA sequences.
It also identifies information about the flexibility of the RNA. Mapping methods are used to build and analyze the folded structure and add important 3D structural considerations.
The model suggests that many biological RNA molecules are optimized by natural selection to fold correctly in the natural context and that stable intermediate RNA secondary structure can persist that anticipates pseudoknot formation.
The model will be published in the online, open-access journal PLoS ONE on September 19.
Source: Public Library of Science