Study of staph shows how bacteria evolve resistance
Antibacterial resistance doesn’t happen overnight. But until recently nobody knew exactly how long it took — or how it happened at all. Now, by studying blood taken from a single patient over a period of months, Rockefeller University researchers have been able to trace how a common strain of bacteria adapted its genes to counteract the antibiotics used to try to kill it, until it finally emerged into the kind of fully resistant microbe that is wreaking havoc in hospitals worldwide. Total elapsed time: 90 days.
This is the first time that such a process has been observed “within” a patient, and the results, published in the May 21 issue of the Proceedings of the National Academy of Sciences sheds light on how such resistance occurs through selective pressure, says the study’s lead investigator, Alexander Tomasz, head of the Laboratory of Microbiology at Rockefeller University.
“What is thrilling is that we got as close as one can to the birthplace of antibiotic resistance in a patient, and now we can study which of the genetic mutations we found are really essential for resistance,” Tomasz says. If the genetic alterations they discovered are common to all known mutated strains of the bacteria — which Tomasz suspects is true — then knowing these genes may help clinicians design ways to block multidrug resistance, he says.
The microbe they isolated is Staphylococcus aureus, which is one of the most frequent causes of a wide range of hospital- and community-acquired infections, and is best known as the cause of toxic shock syndrome. The pathogen has acquired resistance to the majority of available antibiotics, including, recently, vancomycin, which was believed to be the only major agent that could treat it. “It has fantastic adaptive capabilities which have led to the worldwide spread of resistant lineages that are posing serious limits to clinical treatment,” Tomasz says.
But no one has known how such resistance occurs — whether it happens within individual patients, or whether patients with wounds pick up resistant microbes that have somehow infiltrated hospitals.
In this study, Tomasz, along with first author Michael Mwangi, a postdoc in the Tomasz lab, Eric Siggia, head of the Laboratory of Theoretical Condensed Matter Physics, and collaborators from Rockefeller, the Howard Hughes Medical Institute, the U.S. Department of Energy and Cornell University, obtained access to the blood of a patient with congenital heart disease who was treated extensively, but unsuccessfully, with several antibiotics including vancomycin. The team isolated the bacteria from the blood, and then used the whole-genome “shotgun” sequencing method to work out the entire genetic structure of S. aureus as it changed. They sequenced both the initial isolate and the later drug-resistant bacterium.
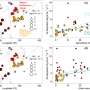
The comparison of the two sequences showed that the resistant bacterium carried 35 mutations in 33 places on its genome and also showed that the mutations showed up in the intermediate isolates in a sequential order in parallel with the gradually increasing resistance to vancomycin. Although initially sensitive to vancomycin, some of the bacteria were probably able to “hide” from the antibiotic in the tissue of the patient’s heart valve, Tomasz says. “The bacteria can bury themselves there and form a wall made of fibrin and platelets, and in that way, microbes in this abscess can selectively adapt to antibiotics in the bloodstream.”
The researchers discovered that as the bacteria acquired resistance to vancomycin, they also became resistant to a new antibiotic, daptomycin, which was thought to be able to treat multidrug-resistant S. aureus. “This is more than we bargained for,” Tomasz says. “The patient wasn’t even exposed to daptomycin, yet the bacteria acquired a resistance to it.” Further testing revealed that one of the mutated loci associated with decreasing vancomycin susceptibility resembled that found from isolates recovered in different regions of the world, raising hopes that these findings will indeed offer a representative model of resistant S. aureus, and may someday lead to new mechanisms for fighting drug-resistant staph.
Citation: Proceedings of the National Academy of Sciences 104(22): 9451-9456 (May 29, 2007)
Source: Rockefeller University