This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Neural network learns how to identify chromatid cohesion defects
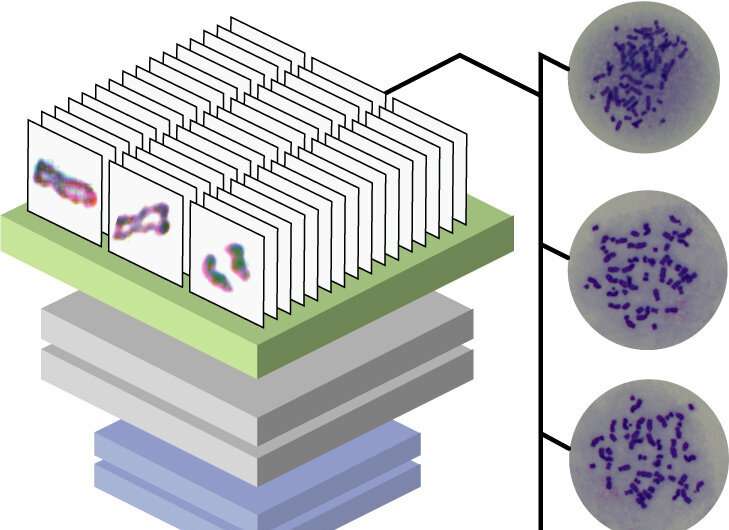
Scientists from Tokyo Metropolitan University have used machine learning to automate the identification of defects in sister chromatid cohesion. They trained a convolutional neural network (CNN) with microscopy images of individual stained chromosomes, identified by researchers as having or not having cohesion defects. After training, it was able to successfully classify 73.1% of new images. Automation promises better statistics, and more insight into the wide range of disorders which cause cohesion defects.
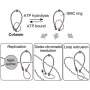
Chromosomes consist of long DNA molecules that contain a portion of our genes. When cells divide, chromosomes need to be copied so that both new cells have all the information they need to function. This is achieved through DNA replication, creating two identical copies known as sister chromatids which are held together by a ring-like protein structure called cohesion. It is vital that these copies be kept together during cell division. Problems with cohesion can lead to the chromatids falling apart, causing serious disruption to the healthy working of cells and organs.
The study of cohesion defects in chromosomes has been largely carried out by researchers observing the chromosomes under the microscope. With a special stain, experienced scientists can tell whether the chromatids are bound in the correct way or not. This kind of classification is vital in the study of chromosomal defects, including the correct functioning of cohesion. However, the entire process is manual. When statistics are required for how many chromosomes are in the correct or incorrect state, the process becomes extremely inefficient, taking vast numbers of man hours by experienced scientists.
Now, a cross-disciplinary team of biologists and machine learning specialists from Tokyo Metropolitan University led by Assistant Professor Takuya Abe, Professor Kiyoshi Nishikawa, Associate Professor Kan Okubo, and Professor Kouji Hirota have combined forces to automate this time-consuming process. They used the same technology that powers facial recognition and machine vision to analyze microscopy images of chromosomes with and without cohesion defects.
In their study published in Scientific Reports, they used a convolutional neural network (CNN), a type of machine learning algorithm particularly well suited to image recognition and trained it on more than 600 images of chromosomes which had been pre-classified into three groups manually by scientists. By the end of the process, new images fed through the algorithm could be classified in the same way as experienced researchers 73.1% of the accuracy. This has the potential to significantly streamline and speed up experiments with chromosome.
The team also used a cell line knocked out a gene known to affect cohesion called CTF18 and analyzed the chromosomes using the trained neural network. The network found significant differences between normal cells and CTF18 knockout cells, indicating that the network, by itself, was capable of picking up genetic problems which impacted cohesion. Though their method currently only recognizes three groups, it can be expanded to different patterns in different species, enabling rapid classification and unprecedentedly precise quantitation of chromosomal defects in a wide range of illnesses.
More information: Daiki Ikemoto et al, Application of neural network-based image analysis to detect sister chromatid cohesion defects, Scientific Reports (2023). DOI: 10.1038/s41598-023-28742-6
Journal information: Scientific Reports
Provided by Tokyo Metropolitan University