Quality control for genetic sequencing
Researchers in the Department of Biosystems Science and Engineering at ETH Zurich in Basel have developed a new method that allows them to record the vast range of antibodies in an individual, genetically in one fell swoop. For example, they can track very precisely how the immune system produces antibodies following a vaccination or an infection. The new genetic method, established by scientists led by Sai Reddy, Professor of Biomolecular Engineering, delivers far more information than the previous decades-old antibody detection techniques.
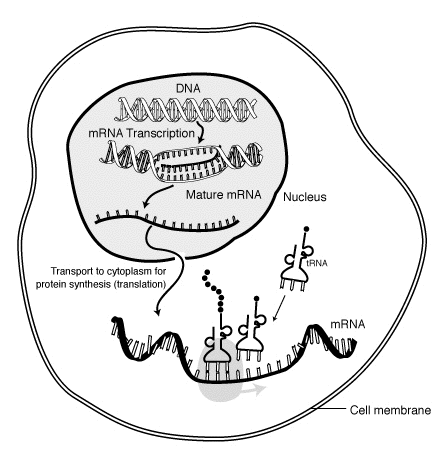
Instead of antibody proteins, the ETH scientist's technique analyses a large number of messenger RNA molecules, which provide instructions to the body's protein production machinery to generate antibodies. Scientists use RNA sequencing to decipher the assembly instructions and to determine their evolution and relative amounts.
A large number of antibodies
"The scientific community has made huge progress in sequencing technology over the last few years. Sequencing has become faster and cheaper, and huge volumes of data can now be processed and analysed," Reddy explains. "Nevertheless, until now, the method was poorly suited to analysis of antibody RNA."
One of the big challenges is that the body contains a huge number of antibodies - with estimates pointing to several billion different variations. However, detecting their differences at the genetic level requires a high degree of sensitivity and precision.
Accuracy a major challenge
To prepare RNA molecules for sequencing, scientists begin by copying their genetic code billions of times. In the process, errors (or mutations) creep in. Until now, it was difficult for scientists to decide whether two slightly different genetic sequences represented two different antibodies, or a single antibody that had mutated during sample preparation.
Furthermore, the sequencing of a mixture of RNA molecules yielded only very imprecise information on the incidence of the respective molecules in the mixture. The reason is that when RNA molecules are copied, as mentioned earlier, not all are duplicated to exactly the same extent.
Over 98 percent of errors avoided
To address this problem, Reddy and his colleagues have supplemented the RNA sequencing process by creating a control system which uses genetic barcodes. This, combined with computer-aided analysis of the sequencing data, has allowed them to massively increase the sequencing accuracy, both regarding artificially introduced mutations and the relative concentration of RNA molecules in the mixture. "In this way, we were able to remove over 98 percent of errors," says Tarik Khan, a postdoc in Reddy's lab.
Specifically, in the new method, each RNA molecule is labelled with a random but unique genetic barcode before amplification. Furthermore, during amplification, the molecules are also tagged with additional unique barcodes in a manner that creates a record of the amplification bias.
Through computer-based analysis of the sequencing data, the scientists can use the barcodes to identify the original antibody RNA molecules (and to differentiate these from molecules that have mutated during the sequencing process). Moreover, using the barcodes and an algorithm, the researchers are able to recover the actual relative amounts of the antibody RNA molecules, thus eliminating copying bias.
Vaccine development and early detection
This new method of antibody sequencing can now be used in immunological research. It is useful, for example, in the development of antibody drugs and vaccines. Reddy is carrying out such work in collaboration with various pharmaceutical companies. "For example, our technique can be used to track precisely how an immune response changes over time, such as in human patients with an HIV infection," says the ETH professor, who was recently awarded one of the coveted Starting Grants from the European Research Council. "Previously, measurements of antibody proteins allowed scientists to discover primarily the very frequent antibodies. However, an immune response always yields a whole range of slightly different antibodies. Sequencing allows us to characterise even the rare ones very accurately - and very quickly."
Furthermore, antibody sequencing can be used to detect small quantities of antibody molecules at an early stage, whereas protein measurement requires a sufficiently high concentration of antibody protein in the blood. Sequencing thus opens up new possibilities in diagnostics; for example, in the early detection of cancer or autoimmunity diseases.
More information: Accurate and predictive antibody repertoire profiling by molecular amplification fingerprinting, dx.doi.org/10.1126/sciadv.1501371
Provided by ETH Zurich