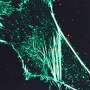
Researchers harness the power of super-resolution microscopy

For centuries, microscopes have been synonymous with science, and today, imaging tools and techniques remain critical to helping researchers zero in on microorganisms and their behavior, providing a window to important clues – and answers – about a host of diseases.
For cellular and molecular biologists like Nathan Roy, Ph.D., a University of Vermont postdoctoral fellow in microbiology and molecular genetics, effective visualization is essential to analyzing proteins – as well as the behavior of lipids, nucleic acids and organelles – within cells. Fluorescence microscopy, which uses a high-intensity light source that stimulates fluorescent elements in a sample, can yield significantly valuable information. However, explains Roy, "the maximum resolution allowed via this technique still has a lower 'diffraction limit,'" which prohibits being able to distinguish truly infinitesimal items. Only recently, says Roy, did independent researchers devise a method to sidestep the diffraction limit.
Enter STORM – Stochastic Optical Reconstruction Microscopy – an imaging system developed at Harvard and then licensed by Nikon. In 2012, the UVM College of Medicine's Microscopy Imaging Center (MIC) installed a Nikon STORM super-resolution microscope, providing investigators with a powerful imaging technology for use in a variety of research endeavors. According to the MIC, this super-resolution imaging technique captures images with a higher resolution than the diffraction limit.
"In some cases, this allows resolution of the equivalent of 10 nanometers, a number previously unheard of using light-based microscopy," Roy says.
UVM College of Medicine faculty members David Warshaw, Ph.D., professor and chair of molecular physiology and biophysics, and Douglas Taatjes, Ph.D., MIC director and professor of pathology, were among the contingent of users that lobbied for UVM's acquisition of this instrument, which, says Roy, "allows the researchers here to stay at the leading edge of current technology."
Nikon N-STORM Super-Resolution Microscope System Features
According to UVM's MIC, the system consists of a Nikon Ti-E TIRF inverted microscope base with laser modules delivering excitation at 405 nanometers (nm), 488 nm, 561 nm, and 647 nm, and a high-sensitivity Andor iXON3 DU897 EMCCD camera. The instrument provides resolution in the fluorescence mode of 20 nm lateral and 50 nm axial. Both fixed and live cell preparations can be imaged, as well as imaging in three dimensions.
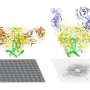
Nikon states that "STORM differs from conventional fluorescence microscopy in that it does not observe all the fluorescently labeled molecules on the sample at the same time, but activates only a very low percentage at any one given time. Repeating this process and acquiring multiple frames in which the molecules are localized with nanometer accuracy results in a final Super-Resolution image."
MIC staff members Taatjes, Nicole Bishop, imaging and analysis specialist in pathology, and Nicole Bouffard, lab research technician in pathology, are well-versed in the system and provide training sessions for researchers.
Research Using STORM Technology
Recently, postdoctoral fellow Adam Sateriale, Ph.D., his faculty mentor Christopher Huston, M.D., associate professor of medicine and infectious disease specialist, and Roy coauthored a paper in PLoS One, titled "SNAP-Tag Technology Optimized for Use in Entamoeba histolytica," which reported results of their research collaboration using UVM's STORM super-resolution microscope.
A waterborne, protozoan bug, Entamoeba histolytica is the primary focus of Huston's research. Entamoeba histolytica infection – also known as amebiasis – is a major cause of parasitic death worldwide, but the pathogenesis of the disease is poorly understood. Huston, Sateriale, and Roy explain in their paper that molecular tools for further studying the disease are limited.
"One problem with studying the parasite Entamoeba histolytica is that there are very few suitable protein tags that can be used to monitor the behavior of proteins in live cells," says Roy.
In an effort to address this issue, Sateriale adapted the SNAP protein tag system – a commercially-available, self-labeling protein tag – to express in Entamoeba. He and Roy teamed up to promote the versatility of his new amoeba SNAP-tag, using STORM to take the first-ever-captured super-resolution images of amoebas.
"By labeling the endoplasmic reticulum (ER), we were able to obtain sub-diffraction images of the amoeba ER – the first of their kind," says Roy. The two researchers believe this new amoeba SNAP-tag should be a useful tool to the worldwide community studying Entamoeba histolytica.
Roy, who completed his Ph.D. in the lab of Markus Thali, Ph.D., professor of microbiology and molecular genetics, specializes in the assembly and transmission process of Human Immunodeficiency Virus type 1 (HIV-1), with emphasis on host factors that can both negatively and positively influence viral replication. He conducted research using STORM for a study he published in the September 2013 Journal of Virology, titled "Clustering and Mobility of HIV-1 Env at Viral Assembly Sites Predict Its Propensity To Induce Cell-Cell Fusion."
For HIV-1 infection to be successful, the virus must fuse its membrane to the target cell membrane, which in turn releases the viral core into the cytoplasm of the target cell.
"This membrane fusion event is mediated by a viral protein called Env," says Roy. "Env has been a major target of vaccines and therapeutics to fight HIV-1 infection, due to the necessity of the membrane fusion step in the viral life cycle."
To gain a deeper understanding of how Env facilitates membrane fusion, Roy, Thali and colleagues Jany Chan, Ph.D., and Marie Lambele, Ph.D., used STORM to study the nanoscale organization and mobility of Env in the plasma membrane of infected cells. They found that the Env was capable of forming ~150 nm clusters, which directly correlated with Env's ability to fuse membranes.
"This suggests that Env clustering is a prerequisite for its function," Roy says.
Using live cell analysis, he and colleagues found that the mobility of Env in the membrane was important for facilitating fusion. While both previously unappreciated properties, the team concludes that the ability of Env to cluster and be mobile are important for its fusion function, and thus HIV-1 spread. Without the Nikon STORM microscope, says Roy, these analyses would not have been possible.
"This new technology has secured our ability to be at the forefront of research that uses imaging tools to study HIV-1 replication," says Roy.
More information: Sateriale A, Roy NH, Huston CD (2013) "SNAP-Tag Technology Optimized for Use in Entamoeba histolytica." PLoS ONE 8(12): e83997. DOI: 10.1371/journal.pone.0083997
"Clustering and Mobility of HIV-1 Env at Viral Assembly Sites Predict Its Propensity To Induce Cell-Cell Fusion." Nathan H. Roy, Jany Chan, Marie Lambelé, and Markus Thali. J. Virol. July 2013 87:13 7516-7525; published ahead of print 1 May 2013, DOI: 10.1128/JVI.00790-13
Journal information: PLoS ONE
Provided by University of Vermont