New method for determining protein structure is 20 times more efficient
(Phys.org) —Research involving scientists from Trinity College Dublin has made a major breakthrough that will streamline the process used to determine the structure of proteins in cell membranes. The development of a new method, and a specialist device to implement the most complex part of the method, should have a major impact on drug-related research.
Over 50% of drugs on the market target cell membrane proteins. It is vital that researchers have high-resolution, 3-D structures of these cell membrane proteins because such 'physiological roadmaps' allow better interpretation of their functional mechanisms. This knowledge can in turn pinpoint weaknesses that can be exploited by drugs specifically designed to act with high selectivity and potency.
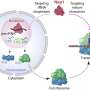
Proteins in cell membranes are also vital for the everyday functioning of complex cellular processes. They act as transporters to ensure that specific molecules enter and leave our cells, as signal interpreters important in decoding messages and initiating responses, and as agents that speed up appropriate responses. As such, it is vital that we know as much about them as possible.
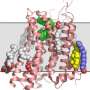
The major challenge facing researchers is the production of large membrane protein crystals. These crystals are used to determine 3-D structure in a complicated process that involves X-rays being fired at them to produce 'diffraction' patterns, which can then be used as 'unique structural fingerprints' of the proteins.
A research group led by Professor of Membrane Structural and Functional Biology at Trinity, Martin Caffrey, developed a high-throughput method for growing membrane protein crystals that makes use of the 'Lipid Cubic Phase' (LCP). The LCP uses a fat-based media to grow these crystals in. Professor Brian Kobilka was awarded his share in the 2012 Nobel Prize in Chemistry, in part for work that made use of the LCP.
Recently, a new method for determining membrane protein structures that uses an X-ray-free electron laser showed great promise. However, it required huge numbers of protein crystals to generate a clear picture of their structure as only 1 in 10,000 was hit in a way that produced useable data. In the breakthrough, Professor Caffrey, as part of a large team of scientists, used the fat-based LCP media in which the protein crystals were grown to jet them across the laser at a relatively slow pace. This slower pace translated into a vastly improved 'hit rate', which in turn provided a more efficient profiling of the protein structure.
In order for the new method to work, the researchers needed to create a device to stream cell membrane protein crystals in the LCP media at a consistent rate. Termed the 'LCP injector', their new device, which resembles a high-tech syringe, is able to adjust the speed of LCP release as required to minimise sample wastage.
Determining the structure of cell membrane proteins has long been seen as a laborious, time consuming, and extremely expensive necessity for medical and therapeutic research. But the new method could change that after the researchers showed it consumed 20 times less protein than the method most commonly used beforehand.
With reference to the research just published in the high-profile journal, Nature Communications, Professor of Membrane Structural and Functional Biology at Trinity, Martin Caffrey, said: "This collaborative work represents a tremendous breakthrough that should greatly improve the efficiency with which research is conducted into determining membrane protein structure."
"The LCP injector shoots the LCP media out at a consistency resembling toothpaste, but at a rate and stream width that must remain highly controlled. Achieving that represented a real challenge and it's very exciting that the final device works so beautifully. Our work will have major applications in the field of drug research and should make it much more efficient and less costly to determine the structure of proteins of interest."
More information: "Lipidic cubic phase injector facilitates membrane protein serial femtosecond crystallography." Uwe Weierstall, et al. Nature Communications 5, Article number: 3309 DOI: 10.1038/ncomms4309. Received 28 November 2013 Accepted 24 January 2014 Published 14 February 2014
Journal information: Nature Communications
Provided by Trinity College Dublin