Characterization of stink bug saliva proteins opens door to controlling pests
Brown marmorated stink bugs cause millions of dollars in crop losses across the United States because of the damage their saliva does to plant tissues. Researchers at Penn State have developed methods to extract the insect saliva and identify the major protein components, which could lead to new pest control approaches.
"Until now, essentially nothing was known about the composition of stink bug saliva, which is surprising given the importance of these insects as pests and the fact that their saliva is the primary cause of feeding injury to plants and crop losses," said Gary Felton, professor and head of the Department of Entomology. "Other than using synthetic pesticides, there have been few alternative approaches to controlling these pests. By identifying the major protein components of saliva, it now may be possible to target the specific factors in saliva that are essential for their feeding and, therefore, design new approaches for controlling stink bugs."
The team reported its results in today's (Feb. 26) issue of PLOS ONE.
According to Felton, stink bugs produce two types of saliva that are required for successful feeding. Watery saliva helps stink bugs to digest their food. Sheath saliva surrounds stink bugs' mouthparts and hardens to prevent spillage of sap during feeding. The hardened "sheath" remains attached to the plant when the insect is finished feeding.
"Unlike a chewing insect, which causes damage by removing plant tissue, stink bugs pierce plant tissue and suck nutrients from the plant," said Michelle Peiffer, research support assistant. "During this process, stink bugs also deposit saliva onto the plant. The interaction between this saliva and the plant is what causes the cosmetic and physiological changes that make crops unmarketable."
To extract the two types of saliva from brown marmorated stink bugs, Felton and Peiffer first collected adult bugs from homes and fields in central Pennsylvania and maintained them in their laboratory.
The researchers chilled the insects on ice. As the insects returned to room temperature, their watery saliva was secreted from the tips of their beaks. The team collected this saliva, processed it and analyzed it for protein content.
To collect sheath saliva, the scientists placed organic grape tomatoes in the cages. After two days of stink bug feeding, they removed the tomatoes from the cages and used forceps to extract the hardened sheaths from the surfaces of the tomatoes. They then processed and analyzed the sheaths for protein content.
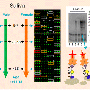
"We found that the watery saliva and the sheath saliva have distinct protein profiles," Felton said. "In other words, we did not find any proteins in common between the two."
Consistent with a role in digestion, the team found that watery saliva contains several digestive proteins, including amylases, proteases and an esterase.
In the sheath saliva, the researchers found peroxidase, suggesting that this protein could be involved in sheath formation. In addition, they found a large number of proteins from tomato.
"These results reveal that the protein composition of the sheath is a mixture of insect- and plant-derived proteins," Felton said. "We used extraordinary precaution to avoid disrupting tomato tissues during the collection of the sheaths, so we do not believe that the composition of tomato proteins in the sheath material is a spurious artifact of our collection methods, but rather it represents the natural coalescing of insect- and plant-derived proteins that occurs during formation of the sheath and subsequent feeding. These initial findings suggest that sheath saliva may elicit a plant self-protection response."
According to the scientists, the methods they developed to extract the saliva and to analyze the proteins should be generally applicable for any species of stink bug.
In the future, the team plans to use a genetic approach to test the function of individual proteins in the saliva to determine their function and essentiality to the feeding process.
"By understanding the specific details of feeding and the damage it causes, researchers can begin to develop targeted control methods for these pests," Peiffer said.
Journal information: PLoS ONE
Provided by Pennsylvania State University