Illinois initiative creates futuristic facility

Through the CompGen initiative, the University's Institute for Genomic Biology and the Coordinated Science Laboratory in the College of Engineering are bringing together top faculty in genomic and computational sciences to create a dynamic team that will develop new technology for genomic breakthroughs.
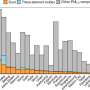
One human genome is made up of about 3 billion nucleotides—enough to fill 130 encyclopedia-sized books that could take nearly 95 years to read, according to the University of Leicester. Four types of nucleotides, represented by A's, T's, C's and G's, create a manual that instructs the cells how to make a human being.
To make sense of this unique manual, geneticists compare numerous sequenced genomes. It's a process akin to trying to find typos or compare words, sentences, paragraphs, and chapters among thousands of books. The new CompGen facility will help researchers analyze trillions of nucleotides to better understand everyone's genetic manual.
"This system will revolutionize genomic research by allowing scientists to reach deeper understandings of highly complex big data sets," said Institute for Genomic Biology director Gene E. Robinson, who initiated the CompGen Initiative with Ravishankar Iyer, Professor of Electrical and Computer Engineering.
Illinois researchers believe this facility, with its state-of-the-art hardware and software coupled with innovative algorithms, will make analyzing DNA more accurate and efficient even as technology advances and researchers are able to sequence larger and larger amounts of data.
"I'm very interested in how to accelerate, how to really speed up, the answers to the many-fold questions geneticists ask," Iyer said. "With a new generation of adaptive processing engines, we can configure them to perform accurate genomic data analysis while accelerating the computation."
Your home computer's central processing unit (CPU) can handle everything you need, but slowly. Through the CompGen initiative, a CPU will be created that does only one thing—analyze genetic data—but does it very quickly.
The data will also be kept secure so that people can choose who sees their genetic manuals.
The Illinois Difference
The innovative processing engines and storage technologies to create this facility are available, or on the horizon. Illinois has the unique multi-disciplinary expertise to make this facility a reality, said Iyer. "I don't believe that anyone else could do it today."
To launch the initiative, Robinson and Iyer invited top biologists, computer scientists and engineers, and bioinformatics specialists at Illinois to participate.
"Normally, you send out an email saying we have this exciting problem and perhaps 10 percent of those people show up," Iyer said. "But here, everybody showed up. They all had an interest in solving this exciting problem."
The CompGen initiative will also promote dialogue between biologists and computer scientists and engineers as they work to develop this new facility. In the past, biologists have struggled to explain their problems in a language that makes sense to computer specialists, while computer specialists have struggled to find solutions that a biologist can understand.
"Each of the two sides is making a big effort to understand the other," said Victor Jongeneel, Director of Bioinformatics and High Performance Biological Computing (HPCBio) at the IGB and a senior research specialist at the National Center for Supercomputing Applications (NCSA). "They know that they need each other."
Visualization
Eventually, the team hopes to incorporate a visualization component to the project that will help researchers visualize genetic data in real time. Giant screens will display real time analytics in the foreground while CompGen's hardware, software, and algorithms work in the background.
It will be the difference between reading about human anatomy in a textbook and seeing human anatomy, and the relationships between body systems, using a 3D application on a tablet.
"Geneticists may not be able to accurately visualize how a particular gene is related to other parts of DNA sequences," said Iyer, also a professor in the Coordinated Science Laboratory and who helped jumpstart the CompGen initiative. "In the past, they couldn't really look and see it, but through new mathematical analysis, we can quantify such multi-way relationships for them."
Support
Already, CompGen has received a $2.6 million grant over 4 years from the National Science Foundation to develop major research instrumentation for this initiative.
Previously, recipients were limited to purchasing equipment off the shelf. Now CompGen has the freedom to create a new machine that is optimized to analyze genomes.
"It's not just about buying cutting-edge hardware," Jongeneel said. "If you just buy hardware, it's an expensive doorstop. The trick is to figure out how to best put this instrument together and how to best adapt it for the tasks at hand."
With financial support from the Office of the Provost and the Office of the Vice Chancellor for Research, CompGen will welcome two new faculty members as well as computer science and engineering students to address these computational problems.
To maximize on CompGen's efforts, Illinois researchers are partnering with more than 15 companies and institutions, including IBM, Abbott Laboratories, Mayo Clinic, Baylor College of Medicine, Microsoft, and the Tata Institute of India, which recently co-hosted a "Computing for Genomics" workshop with Illinois in Bangalore.
"We are extremely excited about the prospects of this new initiative to provide scientists with powerful new tools to address some of the grand challenges in biology related to health, food, energy, and the environment," Robinson said.
More information: compgen.illinois.edu/
Provided by University of Illinois at Urbana-Champaign