New look inside cell nucleus could improve cancer diagnostics
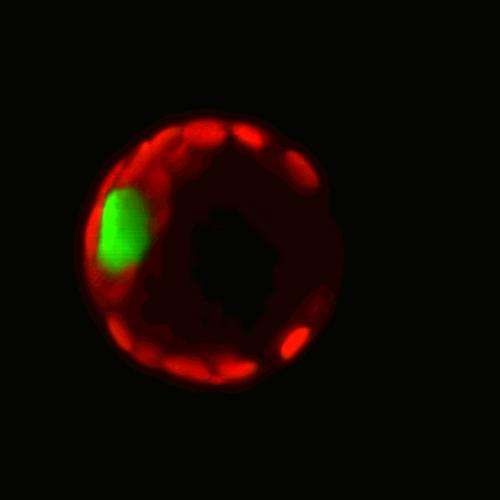
Researchers have successfully isolated and sequenced the entire messenger RNA – the "genetic photocopies" – contained in the nucleus of a single brain cell. This research, published in the journal Proceedings of the National Academy of Sciences, will help researchers better understand how organs function in health and disease and provide another stepping stone toward personalized medicine.
Most cells in animals and plants contain a nucleus, which stores the cell's DNA. Since the DNA never leaves the nucleus, the information stored in the genetic material has to be copied before it can serve in its role as a blueprint for protein synthesis, which happens outside the nucleus. This is accomplished by transcribing sequences of the DNA code into strands of so-called "messenger" RNA. The entirety of messenger RNA at any given time is called the transcriptome; the entirety of DNA inside the nucleus is referred to as the genome.
Transcriptomes tell researchers which genes are being actively transcribed at any given time, and therefore are indicative of a cell's identity and condition.
A team of researchers led by David Galbraith, a professor of plant sciences in the University of Arizona's BIO5 Institute, and Roger Lasken, a professor at the J. Craig Venter Institute in San Diego managed to isolate the transcriptome from the single nucleus of a rat brain cell and decipher the genetic information encoded in it.
By analyzing the transcriptomes of individual cells, researchers hope to better understand the processes that turn, say, a normal cell into a tumor cell. Analyses of single cells have only recently become possible; previously, scientists could only average patterns across many cells, hiding potentially important variation occurring in individual cells.
"The organs and tissues in our body are composed of many different cells, and in order to understand how the organ functions, we have to find out how the individual cells function and disentangle what each of them contributes," Galbraith said. "Further, we need to know how much variation is found in these individual cells."
Previous efforts had shown it was possible to sequence the genetic information encoded in a transcriptome taken from a single cell, but Galbraith's group wanted to know whether it was possible to do the same with just the nucleus since, being copied in the nucleus, that is where the transcriptome originates.

To test their approach, Galbraith and his collaborators isolated the messenger RNA from the nucleus of a mouse brain cell, amplified the number of copies, deciphered the genetic information using high-throughput next generation sequencing, then mapped it back onto the DNA of the known mouse genome sequence.
The method achieved almost the same coverage when applied to single nuclei versus single cells, Galbraith said.
"The correlation between nuclear and cellular RNA was high, and we did not find many transcripts that differed between nucleus and cytoplasm. This tells us we can analyze nuclei and obtain information that is relevant to cells."
Scientists are excited at the idea of studying gene activity at the level of single cells for many reasons. It holds promise for cancer diagnostics, for example, since the much greater degree of resolution available at the single cell level could allow spotting cancerous cells based on their changes in gene activity long before they multiply to manifest clinically as a tumor.
Similarly, comparing the degree of variation in basal gene activity levels within the single cells of different individuals could help physicians identify those with an increased susceptibility to certain types of cancer.
"Before we can do that, it is important to understand how normal cells in tissues differ between each other in terms of their gene activity, and analyzing the transcriptomes from individual cells allows us to do just that."
"Or, when physicians remove a suspicious lesion and isolate and analyze single cells from that lesion, they could build a picture of what is going on in that tissue," Galbraith said. "A subpopulation that is doing something different from the rest could be an early indicator for cancer."
Being able to analyze transcriptomes from nuclei rather than cells has several important advantages in these situations, he explained.
"Brain cells, for example, have very complex interconnections. If we try to isolate the single cells, we have to break these connections and that might well affect the gene activity patterns we want to study."
In addition, separating cells usually involves digestion by enzymes at warm temperatures for an hour or more, resulting in unpredictable changes in the cells that are likely to change gene activities.
Other tissues, such as heart muscle, are very tough, making it impossible to separate the cells from each other without destroying them.
"In both cases, and in most other tissues, getting the nuclei out is easy. You just chop up the samples, and out pop the nuclei," Galbraith said. "Doing this on ice freezes the process of gene expression, which certainly makes one more confident that there are no changes happening that we don't know about."
More information: Rashel V. Grindberg, Joyclyn L. Yee-Greenbaum, Michael J. McConnell, Mark Novotny, Andy L. O'Shaughnessy, Georgina M. Lambert, Marcos J. Araúzo-Bravo, Jun Lee, Max Fishman, Gillian E. Robbins, Xiaoying Lin, Pratap Venepally, Jonathan H. Badger, David W. Galbraith, Fred H. Gage, and Roger S. Lasken. "RNA-sequencing from single nuclei." PNAS 2013 110 (49) 19802-19807; published ahead of print November 18, 2013, DOI: 10.1073/pnas.1319700110
Journal information: Proceedings of the National Academy of Sciences
Provided by University of Arizona