Pushing and shoving: A cost factor in protein synthesis
When cells grow and proliferate, they need to produce large amounts of protein. All this protein is made by ribosomes, therefore rapid growth requires many ribosomes. Because ribosomes are expensive machines for the cell, the cell needs to use them efficiently. In a new study, published in PNAS, a team of researchers from the MPI of Colloids and Interfaces in Potsdam and from universities in Canada, Denmark and the United States investigates how the bacterium Escherichia coli solves this problem and shows that its economic strategy for using ribosomes is close to optimal. This strategy couples increases in the ribosome content of a cell to increase in the speed at which they work: Whenever the cell makes more ribosomes, it also makes them work faster? In addition, the study identified the slow diffusion in the cell as limitation for the speed of ribosomes and as a source for making speed increases costly.
When a company grows, it needs to invest into machines for making larger amounts of its product. The same is true for cells that grow and proliferate. They need to invest their resources into machines such as ribosomes to make protein, and eventually a new cell. In doing so they have to make economic decisions, for example whether it is better to have fewer ribosomes that work faster or more ribosomes that work more slowly. The cell could either put its resources into making many ribosomes or into making them fast. The new study investigates which strategy is used by systematically comparing a theoretical model for proteomic resource allocation with experimental data from Escherichia coli bacteria growing with different nutrient sources and therefore with different growth rates.
The bacteria, it turns out, use a combination of the two strategies: when growth of a cell speeds up, both ribosome content and the speed at which these ribosomes work are increased (but neither as much as it could be if all resources were put into it alone). Is this the best investment strategy that the cell can make? "In our model, it is not optimal but very close", says Stefan Klumpp, group leader at the MPI of Colloids and Interfaces and lead author of the study, "and it is a very simple strategy, so improving it further may come with additional costs."
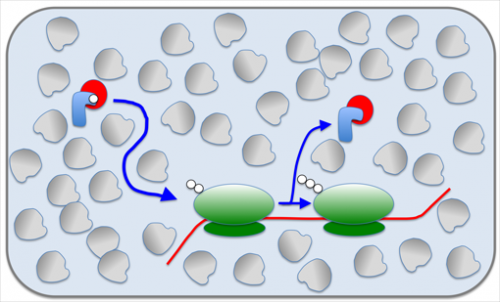
Why is it expensive to make the ribosomes faster? Proteins are chains of different amino acids that are put together in the correct order by the ribosomes. For that the amino acids have to be delivered to the ribosome, which is done by so-called ternary complexes. To make the ribosome work faster, this delivery needs to be faster, but increasing the speed of that delivery faces a physical limit: The interior of a cell is quite crowded with proteins and other molecules, and therefore the big ternary complexes are slow to get through. The only way to make them arrive faster is to have lots of them, so one of them may get to the ribosome in time. Making more ternary complexes in turn requires more ribosomes to make them.
Bacterial ribosomes are the target of many antibiotics, and the study provides a theoretical foundation for understanding the reaction of the cells to these antibiotics. For example, the cells typically compensate the reduced ribosome function partly by making more ribosomes. Similar economic constraints are expected to apply to all kinds of cells, as long as they are rapidly growing and proliferating, so the results may in the future also be applied to higher organisms, in particular cancer cells.
More information: Klumpp, S. et al. Molecular crowding limits translation and cell growth, PNAS, September 30, 2013. DOI: 10.1073/pnas.1310377110
Journal information: Proceedings of the National Academy of Sciences
Provided by Max Planck Society