Biomedical research reveals secrets of cell behavior
(Phys.org) —Knowing virtually everything about how the body's cells make transitions from one state to another – for instance, precisely how particular cells develop into multi-cellular organisms – would be a major jump forward in understanding the basics of what drives biological processes.
Such a leap could open doors to far-reaching advances in medical science, bioengineering and related areas.
An interdisciplinary team of researchers at Arizona State University, with a partner at Imperial College London, report on taking at least a step toward better comprehension of the fundamentals of "cell fate determination" in the prominent research journal Proceedings of the National Academy of Sciences (PNAS).
Cell fate determination relates to the mechanisms by which a cell "decides" in what direction it will go in moving through transitional phases into a final state.
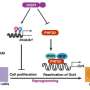
Using mathematical modeling and synthetic biology techniques the team is manufacturing artificial gene networks (a collection of DNA segments in a cell that interact with each other) and introducing them into cells in the laboratory.
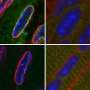
From there, the researchers are able to closely observe through microscopic imaging what is happening with particular cells at their "tipping point," a stage of rest right before they transition into other states.
By learning what takes place at that point, "We can get closer to a fundamental insight about all biology," says biomedical engineer and synthetic biologist Xiao Wang.
Once the mechanisms determining the fate of cells are better understood, Wang says, "We could make gene networks or devices that do what we want them to do," such as create cells that produce medicinal drugs or that kill diseased cells, or create cells that act as sensors to detect environmental hazards.
Wang is an assistant professor in the School of Biological and Health Systems Engineering, one of ASU's Ira A. Fulton Schools of Engineering. He is the senior author of the PNAS paper.
Wang's fellow authors are: biomedical engineering research scientists Min Wu and Xiaohui Li, who work in Wang's lab; electrical engineering graduate student Ri-Qi Su; Ying-Cheng Lai, a professor in ASU's School of Electrical, Computer and Energy Engineering; and synthetic biologist Tom Ellis from Imperial College London.
Their article, "Engineering of regulated stochastic cell fate determination," is available online.
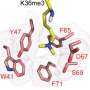
The research team is studying the molecular-level interactions within the DNA sequences of cells, through which the products of one gene affect those of other genes. This helps to trace the lineages of cell development and reveal what drives them in the direction of what kinds of cells they will be in their final states.
Within deeper knowledge of the workings of such processes lays the key to more effectively engineering cells and gene networks.
Wang's team is focused on investigating the intricate properties of gene networks with the goal of learning new ways of regulating the mechanisms behind cell fate determination.
"Our research could be built upon to look at more complicated gene networks and more complex cellular behavior," paving the way for expanding the capabilities of bioengineering to protect and maintain human health, Wang says.
More information: www.pnas.org.ezproxy1.lib.asu. … nt/110/26/10610.full
Journal information: Proceedings of the National Academy of Sciences
Provided by Arizona State University