Why jumping genes don't send us into meltdown
A team of researchers, led by academics at The University of Nottingham, has explained why the so-called 'jumping genes' found in most living organisms don't ultimately kill off their hosts, putting an end to a long-standing scientific mystery.
The study reveals for the first time how the movement and duplication of segments of DNA known as transposons, is regulated. This prevents a genomic meltdown, and instead enables transposons to live in harmony with their hosts—including humans.
Transposons were discovered in the 1940s by Barbara McClintock, who was rewarded in 1983 with the Nobel Prize for Physiology or Medicine. Ancient relics of these 'jumping genes', as they are sometimes called, make up 50 per cent of the DNA in humans. They are characterised as 'jumping' because they can change their position within the genome, thereby creating or reversing mutations. This process, known as DNA transposition, plays a critical role in creating genetic diversity and enabling species to adapt and evolve.
Transposons don't just jump from one location to another; they usually leave behind a copy of themselves at their original location. Left unchecked, this would lead to an exponential increase in their numbers. Exponential growth is always unsustainable, and in the case of transposons they would quickly kill their host.
Because this doesn't happen, clearly some form of regulation is taking place within the genome. For a long time, scientists have understood that an enzyme called transposase, which is critical to the whole transposition process, also apparently brings it under control.
How this actually occurs has, however, remained a mystery—until now. For the first time, the new study, carried out by researchers at The University of Nottingham, The University of Cambridge and the Fred Hutchinson Cancer Centre in Seattle, successfully identified the mechanism through which DNA transposition is regulated.
Ronald Chalmers, Professor of Molecular and Cellular Biology at The University of Nottingham, said "A successful parasite is not fatal to its host but lives in harmony with it. It was while doing some biochemistry research that we stumbled upon the solution. It was so simple that initially it was hard to appreciate the brilliance of it. It was a real forehead-slapping moment and seems a wonder that it was not discovered years ago."
"In theory, transposons should just keep on increasing and kill us," said Dr Karen Lipkow, from the Department of Biochemistry, University of Cambridge, and the Babraham Institute. "What we have identified is an ingenious mechanism which prevents this from happening by conferring autoregulation. The process is very simple, but it explains so much."
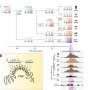
A biochemical analysis of a mariner-family transposon from the human genome gave the first indication of the mechanism. The team then expanded this concept using a computer model, which allowed them to simulate events on an evolutionary timescale, before carrying out further biochemical experiments.
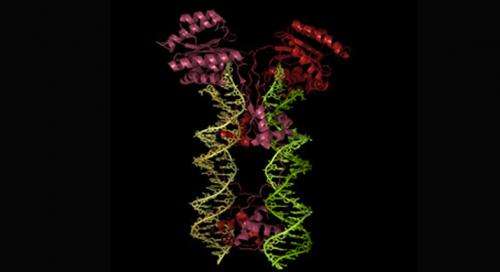
Transposase is important to the genetic 'jump' of transposons because it catalyses the whole process. It does this by binding to specific sites at either end of the transposon, and then bringing these together to create a nucleoprotein complex that effectively performs a molecular 'cut and paste, moving the transposon DNA to a new location.
The study found that once a certain number of copies of the transposon have been created, the transposase concentration rises above a critical threshold and begins to saturate its own binding sites. As clusters of the enzyme compete for binding, they interfere with each other and the transposition process is halted.
Using the computer model, the researchers were able to show that doubling the transposon copy number halves the rate of transposition. This is a crucial insight as it reveals an underlying self-righting mechanism for homeostasis. In a nutshell, it makes the transposon's genetic burden on the host constant and predictable, and brings an uneasy harmony to the relationship.
More information: elife.elifesciences.org/content/2/e00668
Provided by University of Nottingham