Finding Nematostella: An ancient sea creature
A study of tentacle-formation in a sea anemone shows how epithelial cells form elongated structures and puts the spotlight on a new model organism.
There's a new actor on the embryology stage: the starlet sea anemone Nematostella vectensis. Its career is being launched in part by Stowers Institute for Medical Research Associate Investigator Matt Gibson, Ph.D., who is giving it equal billing with what has been his laboratory's leading player, the fruit fly Drosophila melanogaster.
Gibson's lab investigates the cellular and molecular mechanisms used by cells to assemble into layers or clusters during embryogenesis. Those tissues, comprised of densely packed cells known as epithelial cells, shape the body not only of simple creatures but also of mammals, where they line every body cavity from lung to intestine and form hormone- and milk-secreting glands. Unfortunately these cells have a dark side too- over 80% of human cancers, carcinomas, are of epithelial origin.
The Gibson lab has historically used the genetic powerhouse Drosophila to investigate the control of epithelial cell shape and proliferation during wing, leg and eye development. Breaking with tradition, their new study published in the May 15th, 2013 issue of Development, explains how developing sea anemone larvae construct an even more basic epithelial appendage, the tentacle. The paper charts how epithelial cell shape changes drive tentacle development and is also the first to identify candidate genes driving those changes. Most of all, by putting a new model organism representing one of the simplest animals center stage, the study illuminates some of the most fundamental principles animals use to construct a body.
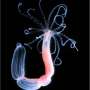
Lacking even left-right symmetry, sea anemones are evolutionarily ancient. But during embryogenesis their larvae compensate for an uninspiring torso by sprouting tentacles from thickened epithelial buds surrounding their mouth. "Nematostella's body is basically a bag of epithelium," says Gibson. "And that simplicity makes it a great system for determining how epithelial cells act collectively to shape an appendage. Taking advantage of this fast, easy and cheap experimental system, we can quickly answer questions that give us deep insight into a process, at both the mechanistic and evolutionary levels."
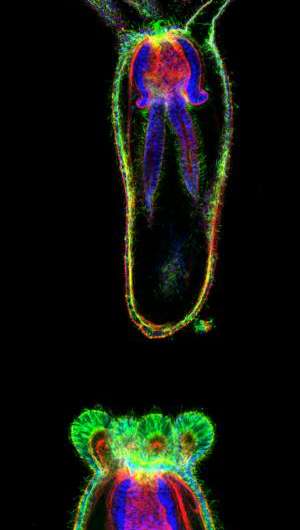
The all-Stowers study, led by first author Ashleigh Fritz, a graduate student at the University of Kansas School of Medicine working in the Gibson lab, began by imaging Nematostella larvae at the cellular level before, during, and immediately after "juvenile" tentacles sprang from their body. Freshly hatched Nematostella larvae are under intense pressure to get their tentacles up and running, as they use them to pull food toward their mouths. The question was, what kind of cellular reshuffling drove these survival-dependent changes in morphology?
"We thought tentacle outgrowth might be driven by cell proliferation," says Fritz, noting that some of Nematostella's freshwater cousins sprout appendages by constant cell division. "Instead, we observed that cells begin thickened and then thin out as tentacles elongate." In other words, the process was driven not by cell duplication along a "tentacle axis" but rather by stretching a stockpile of cells.
Embryologists call the embryonic thickening of epithelial cells that provides raw material for a mature structure a placode. "Placodes have appeared over and over throughout evolution," says Gibson, noting that placodes give rise to wings or eyes in flies and feathers and teeth in vertebrates. "Discovering that placodes are also utilized in animals as seemingly primitive as Nematostella shows how fundamental this strategy is in evolution."
The group also showed that activation of a cellular receptor known as Notch was mandatory for tentacles to emerge from a placode. Newly hatched Nematostella larvae swimming in lab seawater laced with a drug that blocks Notch receptor activity failed to sprout tentacles.
The researchers also constructed microarrays from tissue isolated at early, mid, and late stages of tentacle extension, allowing global comparison of the collection of mRNAs, or the "transcriptome", at each stage. That effort, driven by Stowers Research Advisor Chris Seidel, Ph.D., and Ariel Paulson of the Stowers Computational Biology Core, is an obligatory step in pioneering any new model organism.
"Transcriptome analysis led us to identify novel tentacle markers," says Fritz, referring to molecular probes used to define a particular cell type. "Also gene expression patterns that we and others have identified allowed us to construct the first-ever molecular model of how tentacles are patterned."
In short, the study not only suggests universal principles underlying sculpting of epithelial structures from a placode, but also provides investigators with a toolkit to test whether specific genes drive the process.
An added bonus is that in 2007 a consortium of researchers sequenced the Nematostella genome and reported it to be more "human-like" in size and structure than that of Drosophila or another widely used model system, the nematode C. elegans. As a result, Gibson thinks that for many key questions, Nematostella may represent a better laboratory model than either.
"The common ancestor of sea anemones, flies, and humans likely had a surprisingly complex genome," he says, explaining that over millions of years of evolution flies and worms might have lost some genomic complexity. "As a result, these seemingly simple animals share some key genomic characteristics with humans and other vertebrates."
The Gibson laboratory continues to use both flies and sea anemones to ask how epithelial proliferation is controlled and why epithelial placode formation is so prevalent in developing embryos. Their next task is to develop molecular approaches to test how specific genes govern Nematostella embryogenesis. "Right now we are actively working on experimental tools, including techniques to knockout, edit or overexpress genes in Nematostella," says Gibson. "This paper opens up new ground and lays foundation for a next round of more deeply mechanistic studies."
Journal information: Development
Provided by Stowers Institute for Medical Research