April 23, 2013 report
Researchers map primate networks to predict pandemics
(Phys.org) —Most emerging infectious diseases (EIDs) that affect humans originated in animals. However, epidemiologists have been unable to identify the sources of zoonotic diseases until after they have already infected humans. Using network science, Jose M. Gomez and his colleagues at the University of Granada, the University of Valencia and Harvard University have developed a method for predicting which species are most likely to transmit EIDs to humans. Their research appears in the Proceedings of the National Academy of Sciences.
The study of networks as a way of predicting disease transmission is not new. Epidemiologists study social networks to understand the connections between different people. They use this information to predict how infectious diseases will spread.
Environmental scientists map networks between species to illustrate how they interact with each other.
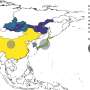
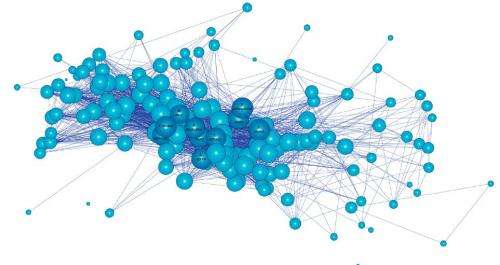
Gomez and his team combined these uses of network science, mapping a network that linked pathogen-carrying species by the parasites they carry to study inter-species disease transmission. They examined records of 140 non-human primate hosts harboring 300 parasite species, linking species that shared the same parasites. The researchers focused on non-human primates because they are closely related to humans and share habitats with humans; therefore they are the most likely to harbor parasites that will infect us.
Species that shared the most parasites with the most other species were at the center of the network. These species tended to have wide geographic ranges and to live in large, tightly packed social groups, conditions that increase the likelihood of contact with other species. These "superspreaders" were the most likely to harbor pathogens resembling those found in humans and to host previously identified EIDs.
As expected, apes, our closest relatives and the source of infectious diseases such as HIV, malaria and yellow fever, had central positions in the network. However, baboons, macaques, squirrel monkeys, howler monkeys and vervet monkeys also appeared at the network center.
Gomez and his team suggest that scientists could help ward off the next pandemic by keeping an eye on species that live in dense social groups and cover wide geographic ranges. They could test superspreaders for viruses that mutate at high rates, increasing the chance that they will infect other species.
The researchers say that this methodology need not be restricted to primates. Other animals, including birds, pigs, bats and rodents, have also been the source of EIDs. A network approach could help identify all species that pose a high risk of transmitting diseases to humans.
More information: Centrality in primate–parasite networks reveals the potential for the transmission of emerging infectious diseases to humans, PNAS, Published online before print April 22, 2013, doi: 10.1073/pnas.1220716110
Abstract
Most emerging infectious diseases (EIDs) in humans have arisen from animals. Identifying high-risk hosts is therefore vital for the control and surveillance of these diseases. Viewing hosts as connected through the parasites they share, we use network tools to investigate predictors of parasitism and sources of future EIDs. We generated host–parasite networks that link hosts when they share a parasite, using nonhuman primates as a model system because—owing to their phylogenetic proximity and ecological overlap with humans—they are an important source of EIDs to humans. We then tested whether centrality in the network of host species—a measurement of the importance of a given node (i.e., host species) in the network—is associated with that host serving as a potential EID source. We found that centrality covaries with key predictors of parasitism, such as population density and geographic range size. Importantly, we also found that primate species having higher values of centrality in the primate–parasite network harbored more parasites identified as EIDs in humans and had parasite communities more similar to those found in humans. These relationships were robust to the use of different centrality metrics and to multiple ways of controlling for variation in how well each species has been studied (i.e., sampling effort). Centrality may therefore estimate the role of a host as a source of EIDs to humans in other multispecific host–parasite networks.
Journal information: Proceedings of the National Academy of Sciences
© 2013 Phys.org