Explainer: What is a gene?
There's a very confusing exchange in Lewis Carroll's Through the Looking Glass: "When I use a word," Humpty Dumpty said, in rather a scornful tone, "it means just what I choose it to mean—neither more nor less."
When people use the word "gene" it's also important to know what they intend it to mean. The meaning may depend on whether one is talking about carrying a gene, expressing a gene, transferring a gene or discussing how many genes we have.
One reason the definition is so confusing is that the term was coined in 1909, before we really knew what a gene was. And the effects of genes – inherited characteristics – were observed before we understood genes.
As our knowledge has advanced, the definition of the word gene has evolved; and with all the information from the new ENCODE project, the definition needs updating again. For an excellent academic summary of the current definition see the recent paper and poster in the journal Genome Research.
The molecular basis of inheritance
The Austrian monk Gregor Mendel carried out the first genetics in the 1860s and showed that characteristics were inherited.
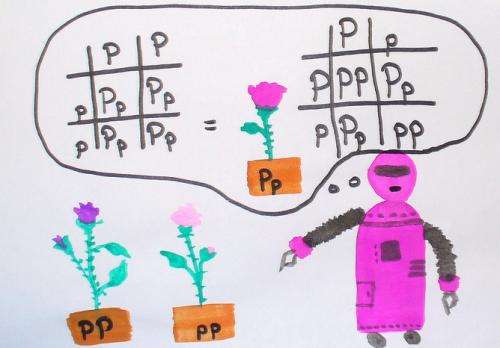
We have always known that pea seeds grow into pea plants, not into kangaroos. What's more, plants with red flowers usually have offspring that have red flowers: children resemble their parents.
Mendel showed that crossing a red pea with a white pea could give rise to peas that were not pink but were either white or red.
We miss this point sometimes because we all have features from our two parents, and many features seem to blend. But Mendel showed that distinct characteristics could be inherited intact and we can think of these as each being encoded by a gene.
But Mendel never used the word "gene". Nor did Darwin. The word gene was first used in 1909 by the Danish botanist Wilhelm Johnannsen to refer to "determiners which are present [in the gametes] … [by which] many characteristics of the organism are specified".
Later it was found that, whatever material carried these characteristics, it was linear, like string. In 1915 the American geneticist Thomas Morgan found some genes tended to be co-inherited (flies might co-inherit short wings and red eyes together from one parent, more often than short wings and short legs).
He deduced this might mean certain genes were close together, much like beads on a string.
The idea that the genetic material was linear was born. But we still didn't know what it was.
In the 1940s the American physician Oswald Avery showed that an enzyme that chews up DNA, DNase, could destroy genes.
We finally knew that the genetic material was DNA.
In 1953 James Watson and Francis Crick, along with Maurice Wilkins, using data from Rosalind Franklin, showed DNA was found in the form of a double helix. The fact it was double, with two matching strands, suggested how it could be replicated.
First definitions of a gene
But what precisely was a gene? Crick explained how DNA could be "transcribed" into RNA (ribonucleic acid) and RNA could be "translated" into protein. Think of a protein as a biological tool that does something – i.e. the haemoglobin that carries oxygen in your blood.
This gave us our first solid definition:
A gene is a stretch of DNA that encodes a piece of RNA that encodes a chain of protein.
The technical details are complex but let's imagine how you might make a metal axe, or many axes.
Picture a segment of DNA bundled in the precise shape of an axe-head. Consider the RNA nestles in and forms the impression of an axe-head – so it's now like a mould or cast.
The RNA travels out of the DNA storage room – the nucleus – and you pour in molten iron. It hardens and out comes an axe-head. You would have another mould for the metal handle.
The axe-head and handle then bounce around in the cell, find each other and self-assemble. Post-translational modifications, akin to sharpening, can be done by other machines in the cell.
If we mould a lot of axes then we say the gene is expressed at high levels; if there are few or no axes, the gene is expressed at a low level, or is silent.
We can make use of the axe analogy in another way. One definition of a gene is a region that makes a protein tool. But there are many DNA genes that make RNA and the process stops there.
Similarly, the RNA for an axe-head – or one like it – might make a perfect holder for an axe: it doesn't need to go on to make the axe itself.
This gene would produce what is called a "non-coding RNA" – an RNA that has a function in itself and doesn't need to encode a protein.
Early life-forms probably used RNA and got by without proteins or DNA. Our oldest cellular tools – tRNA and rRNA – which work on the assembly line, making proteins, are never themselves translated into protein.
Most interestingly, recent work, such as the ENCODE project, suggests we have underestimated the number of non-coding RNAs. One problem, though, is that there also appears to be noise RNA that probably does little harm but no good either, so not every RNA will be functional. Not every segment of DNA that encodes an RNA is a gene.
At this stage it is important to point out that, there are no actual casts or molten iron but instead strings of Lego-like blocks of different shapes. A section of the DNA blocks is read into RNA blocks.
The RNA blocks are read into 20 different protein-building blocks that fold up according to their shape to make, in this instance, an axe handle perhaps.
The axe doesn't actually resemble the DNA or the RNA in shape at all; the sequence of Lego blocks is dictated by the sequence in the DNA, via a special code – called the genetic code.
A definition at last
But now we have a definition for a gene:
Genes are stretches of DNA that have the potential to create a tool or a characteristic – such as red colour in the pea flower.
The outcome is called the "phenotype," and our "genotype" (our genetic material) plus environmental inputs create our phenotype. The Human Genome Nomenclature Organization defines a gene as "a DNA segment that contributes to phenotype/function".
A gene is a linear section of DNA – of a chromosome – that contributes some function to the organism. There are many genes on each human chromosome – thousands.
There are also spacer regions between genes and even within genes (introns) that may or may not do anything.
Some do – the major control region of the gene (the promoter) sits just upstream or around the start point of the gene; but there are also enhancer and silencer elements that can be positioned at very great distances along the chromosome and regulate the level of expression.
It is not clear whether or not to include the control regions as part of the gene. Strictly speaking, the gene is usually only the "coding part" – the mould – but mutations in the control regions can be just as damaging as those in the mould itself.
So one good definition of a gene is the entire DNA region that is necessary for the synthesis of a functional RNA or protein.
At first, each gene was thought to produce one protein tool, but we can use our analogy of the axe to explain how one gene can produce more than one protein or tool.
The axe handle gene might be "spliced" – a process where bits are cut out of the RNA transcript before it is translated into protein. In this way we might produce a short handle to make a tomahawk or hatchet instead of a full-sized axe.
The amount of alternative splicing in humans is extensive and typically several gene products are made from each gene.
The suggested post-ENCODE definition of a gene is:
A union of genomic sequences encoding a coherent set of potentially overlapping functional products.
Why would we have evolved a gene for cancer?
We can now also explain what it means for a plant to carry the gene for red flowers – it may mean that the plant has a stretch of DNA that encodes an enzyme (a protein tool that catalyses chemical reactions) to make a red coloured pigment.
But is there a gene for white flowers? There may just be a mistake in the red flower gene so that the enzyme no longer functions, so the flowers have no colour.
This also explains the confusion between describing the gene in terms of the tool it makes or the ultimate effect of that tool.
What does carrying the gene for breast cancer mean? It doesn't mean a special gene has evolved and is out there with the function of causing breast cancer.
It means a gene involved in limiting cellular doubling in breast tissue or in DNA maintenance is mutated and no longer functions. So the probability of a cancer growing is increased. The gene predisposes the carrier to cancer – it doesn't cause it.
The gene for hemophilia is not there to cause bleeding: it's a gene that, when mutated, results in a defective clotting factor and bleeding is the result.
There are several genes for breast cancer and there are two common genes for hemophilia. Just as mutating the axe-head gene or the axe-handle gene would cause the axe to fail, many biological proteins work together or in pathways, and breaking any link in the chain can have serious outcomes.
The most confusing thing is that the "gene for breast cancer" may have a very indirect relationship to the biology of the breast.
If people were planets and one had a mutation in its axe handle gene, a molecular biologist would observe that there were no functional axes on the planet, but a geneticist would have first noticed that the world was covered in trees.
The gene wouldn't be called the axe gene, but would first be noticed as the gene for making forests. It would only be later that someone would map the gene, clone it and find out what it encoded and how its product functioned.
How many genes do we have?
We still don't know how many genes we have for certain. A famous sweepstake was carried out when the human genome – all our DNA – was first sequenced, and estimates were as high as 100,000. But we now think the number is much fewer.
One can spot many genes by computer since they have certain key features – an RNA is read from them and the genetic code translates into a protein of reasonable length. But it's hard to identify short genes and functional non-coding RNA genes.
There are probably about 20,000 genes encoding proteins and perhaps as many encoding functional RNAs. We don't know the precise number because it is very hard to be sure which segments of DNA are read into functional products. We won't know that unless they are mutated.
And that's an experiment no-one will be doing on humans, although, as our information on existing human populations and other species increases, we are sure to improve our knowledge of the vast genomic wonderland and to discover new genes we didn't know were there.
Journal information: Genome Research
Source: The Conversation
This story is published courtesy of The Conversation (under Creative Commons-Attribution/No derivatives).
















