New system to improve DNA sequencing

(Phys.org) —A sensing system developed at Cambridge is being commercialised in the UK for use in rapid, low-cost DNA sequencing, which would make the prediction and diagnosis of disease more efficient, and individualised treatment more affordable.
Dr Ulrich Keyser of the University's Cavendish Laboratory, along with PhD student Nick Bell and other colleagues, has developed a system which combines a solid-state nanopore with a technique known as DNA origami, for use in DNA sequencing, protein sensing and other applications. The technology has been licensed for development and commercialisation to UK-based company Oxford Nanopore, which is developing portable, low-cost DNA analysis sequencing devices.
Nanopore technology has the potential to revolutionise DNA sequencing and the analysis of a range of other biological molecules, providing dramatic improvements in power, cost and speed over current methods.
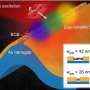
A nanopore is an extremely small hole - between one and 100 nanometres in diameter – typically contained in a membrane between two chambers containing a salt solution and the molecule of interest. When the molecules pass through the nanopores, they disrupt an ionic current through the nanopore and this difference in electrical signals allows researchers to determine certain properties of those molecules.
Over the past decade, researchers have been investigating various methods of constructing nanopores in order to improve accuracy and reliability. A key part of this is the ability to finely control the shape and surface chemistry of the nanopores, which would maximise sensitivity and facilitate the identification of a wider range of molecules.
Currently, there are two main types of nanopores in use: solid state nanopores constructed by fabricating tiny holes in silicon or graphene with electron beam equipment; and biological nanopores made by inserting pore-forming proteins into a biological membrane such as a lipid bilayer.
Biological nanopores are cheap and easy to manufacture in large quantities of identical pores. It is possible through genetic engineering to define their structure at the atomic level, varying the pores for the analysis of different target molecules. However, they are only suitable for a limited range of applications, and may be replaced over time by solid-state nanopores. At present, solid-state nanopores are difficult to manufacture and are not as sensitive as biological nanopores, as it is difficult to position specific chemical groups on the surface.
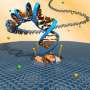
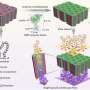
In collaboration with researchers at Ludwig Maximilian University in Munich, Dr Keyser and his team have developed a hybrid nanopore which combines a solid-state material, such as silicon or graphene, and DNA origami - small, well-controlled shapes made of DNA.
"The DNA origami structures can be formed into any shape, allowing highly accurate control of the size and shape of the pore, so that only molecules of a certain shape can pass through," says Dr Keyser. "This level of control allows for far more detailed analysis of the molecule, which is particularly important for applications such as phenotyping or gene sequencing."
Since complementary sequences of DNA can bind to one another, the origami structures can be customised so that functional groups, fluorescent compounds and other molecular adapters can be added to the DNA strands with sub-nanometre precision, improving sensitivity and reliability. Additionally, hundreds of billions of self-assembling origami structures can be produced at the same time, with yields of up to 90 per cent.
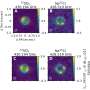
Recent research by the team, published in the journal Lab on a Chip, has shown that up to 16 measurements can be taken simultaneously, allowing for much higher data throughput and screening of different DNA origami structures.
More information: pubs.rsc.org/en/content/articl … g/2013/lc/c3lc50069a
Journal information: Lab on a Chip
Provided by University of Cambridge