March 28, 2013 report
Which came first the head or the brain?

(Phys.org) —A fundamental question in the evolution of animal body plans, is where did the head come from? In animals with a clear axis of right-left symmetry, the bilaterians, the head is where the brain is, at the anterior pole of the body. Little is known about the possible ancestor of bilaterians. Fortunately their sister group from that same progenitor, the cnidarians, can be studied in parallel today to give some clues. Cnidarians are creatures like jellyfish, hydra, and sea anemone which possess rudimentary nerve nets, but no clear brain. They all have just a single orifice to the external world, which basically does it all. In a recent paper published in PLOS Biology, researchers from the University of Bergen in Norway compared gene expression patterns in sea anemone (Nematostella vectensis, Nv) with that from a variety of bilaterian animals. They found that the head-forming region of bilaterians is actually derived from the aboral, the opposite-oral, side of the ancestral body plan.
Pioneering developmental biologist Lewis Wolpert, is often credited with having observed: "It is not birth, marriage, or death, but gastrulation which is truly the most important time in your life." Almost all animals undergo a similar gastrulation process early in their development. The point where the cells first invaginate during gastrulation, the blastopore, uniquely defines an embryonic axis. After this stage however, all bets are off—attempts to define phyla according to hardline criteria, like blastopore = anus, are invariably met by counterexample where it instead becomes the mouth. Gene expression, while not always constrained into single contiguous areas, therefore provides a baggage-free way to assign homology across species.
Wolpert's concept of positional information in development has been largely vindicated by the discovery of hox gene codes in a wide variety of animals. While hox genes are the critical regulators of axial patterning, in most bilaterians they are not expressed in the anterior head-forming region. The researchers focused instead on the genes six3 and FoxQ2, transcription factors which have been shown to regulate anterior-posterior development. Six3 knockouts in mice, for example, fail to develop a forebrain. In humans six3 regulates forebrain and eye development.
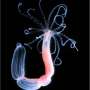
Sea anemone, like Nematosella, are curious creatures. As larvae they swim about with their aboral pole forward. As adults they plunge this region into the sea floor, and permanently anchor themselves in. Their bodies then undergo various changes but their oral pole remains intact for feeding. By using knockdown and rescue experiments in Nemostella, the researchers were able to show that six3 is required for the development of the aboral region, and the expression of further regulatory genes. This suggests that the region distal from the cnidarian mouth parallels development of the bilaterian head.
The researchers also looked at the expression of the forkhead domain protein foxQ2, which functions downstream of six3. Forkhead box genes are an important class of transcription factors which frequently lack the signature homeodomains and zinc-finger regions common to other transcription factors. Instead they have a unique DNA-binding region that has the shape of a winged helix. The forkhead gene, fox2p, in humans has recently garnered a lot of media attention for its apparent role in neural development, and in even more esoteric functions like speech development.
FoxQ2 is known to be a well-conserved marker for the most anterior tip of a variety of bilaterians including sea urchines, drosophila, and cephalochordates. The researchers established that before gastrulation in cnidarians, foxQ2a was expressed in the aboral pole, and in a small number cells resembling neurons. Afterwards the expression of this "ring gene" was excluded from a central spot.
In conclusion, the expression of genes for anemone head development, away from the mouth region, suggests that head development came first and was a separate event from mouth development. Secondarily, the head and a coalescing brain appear to have merged to become a centralized control center.
More information: Sinigaglia C, Busengdal H, Leclère L, Technau U, Rentzsch F (2013) The Bilaterian Head Patterning Gene six3/6 Controls Aboral Domain Development in a Cnidarian. PLoS Biol 11(2): e1001488. doi:10.1371/journal.pbio.1001488
Abstract
The origin of the bilaterian head is a fundamental question for the evolution of animal body plans. The head of bilaterians develops at the anterior end of their primary body axis and is the site where the brain is located. Cnidarians, the sister group to bilaterians, lack brain-like structures and it is not clear whether the oral, the aboral, or none of the ends of the cnidarian primary body axis corresponds to the anterior domain of bilaterians. In order to understand the evolutionary origin of head development, we analysed the function of conserved genetic regulators of bilaterian anterior development in the sea anemone Nematostella vectensis. We show that orthologs of the bilaterian anterior developmental genes six3/6, foxQ2, and irx have dynamic expression patterns in the aboral region of Nematostella. Functional analyses reveal that NvSix3/6 acts upstream of NvFoxQ2a as a key regulator of the development of a broad aboral territory in Nematostella. NvSix3/6 initiates an autoregulatory feedback loop involving positive and negative regulators of FGF signalling, which subsequently results in the downregulation of NvSix3/6 and NvFoxQ2a in a small domain at the aboral pole, from which the apical organ develops. We show that signalling by NvFGFa1 is specifically required for the development of the apical organ, whereas NvSix3/6 has an earlier and broader function in the specification of the aboral territory. Our functional and gene expression data suggest that the head-forming region of bilaterians is derived from the aboral domain of the cnidarian-bilaterian ancestor.
Synopsys: www.plosbiology.org/article/in … journal.pbio.1001484
Journal information: PLoS Biology
© 2013 Phys.org