Gap geometry grasped: New algorithm could help understand structure of liquids, how they flow through porous media
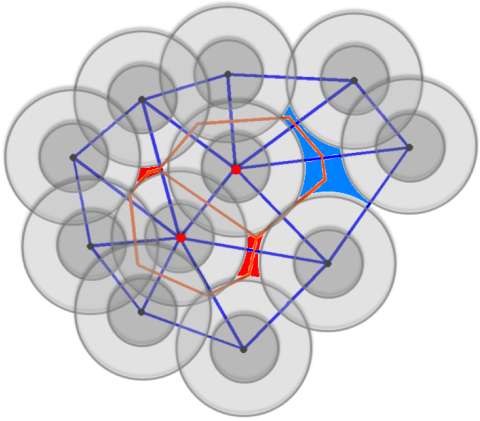
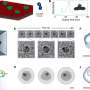
Theoretical physicist Moumita Maiti and colleagues at the Jawaharlal Nehru Centre for Advanced Scientific Research in Bangalore, India, have now implemented an algorithm for analysing void space in sphere packing, where the spheres need not all be the same size. This method, about to be published in European Physical Journal E, could be applied to analyse the geometry of liquids present between multi-sized spheres that are akin to a model for porous material. This provides a tool for studying the flow of such fluids through porous material. More importantly, it can also be used to study the packing geometry of proteins.
There have been several previous attempts to calculate the volume and the surface area of packing of spheres. But few methods have taken into account the connectivity of empty space between spheres, which matters, for example, when detecting buried cavities in proteins.
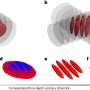
To remedy this issue, the authors have relied on a programme capable of performing a very detailed study of the size distribution of the free volumes of individual spheres-that is, the volume swept by the centre of the sphere without overlapping with any of the other spheres-in jammed sphere packing.. It also makes it possible to calculate the exact volumes and surface areas of cavities by detecting the disconnected components of cavities.
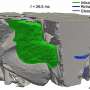
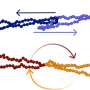
The team applied this method to the analysis of protein structures. This led them to compute various key quantities such as the distribution of sizes of buried cavities and pockets between spheres, the matching of areas accessible to solvent in which protein are found with the corresponding volumes and the composition of residues lining cavities.
Ultimately, the authors are planning to prepare this algorithm for distribution as open source software.
More information: M. Maiti, A. Laxminarayanan and S. Sastry (2013), Characterization of Void Space in Polydisperse Sphere Packings: Applications to hard sphere packings and to protein structure analysis, European Physical Journal E, DOI 10.1140/epje/i2013-13005-4
Journal information: European Physical Journal E
Provided by Springer