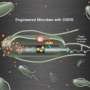
Modified bacteria turn waste into fat for fuel

"Green" chemistry developed at Rice University is at the center of a new government effort to turn plant waste into fatty acids, and then into fuel.
The Rice lab of bioengineer Ka-Yiu San is part of a recently announced $25 million United States Department of Agriculture project to develop a new generation of renewable energy and bio-based products from switchgrass and forestry residues and from a new hybrid of sorghum being developed at Texas A&M University.
Patent-pending fermentation processes created by San and his colleagues use genetically modified E. coli bacteria to produce fatty acids from hydrolysates. The sugary, carbon-rich hydrolysate is extracted from cellulose, the tough, inedible part of plants that is usually thrown away. San said his lab already gets an 80-to-90 percent yield of fatty acids from model sugars and hopes to improve that over the next few years.
"Adding another 1 or 2 percent doesn't seem like much," said San, based at Rice's BioScience Research Collaborative. "But when you're talking about making several million tons per year, it's huge."
The target products are synthetic diesel and lubricants, according to Ceramatec Inc., a Utah-based company that proposed the project and would produce hydrocarbons from fatty acids that could then be processed by petroleum refineries.
There are two ways to make fuel (from biomass)," said Mukund Karanjikar, innovation manager at Technology Holding LLC, which is administering the project. "You either make alcohols, or you make petroleum-like fuels that can go into current infrastructure. Our program is for infrastructure-compatible transportation fuels.
"There aren't many ways to go from sugars to a diesel-like compound," he said. "The best way is to make fatty acids from the sugars microbially, as many labs have tried to do. But the Rice University process is definitely the winner."
Postdoctoral researchers Xiujun Zhang and Mai Li have been nudging their bacteria toward efficient production of fatty acids for four years, San said. Zhang is responsible for the development of enzymes in E. coli that promote the efficient formation of free fatty acids, while Li, now at GlycosBio, worked to build microbial host strains for high-yield production.
"They have been instrumental to this project," he said. "In four years, with two postdocs, we beat everybody, even groups with dozens of members."
San said the researchers screened hundreds of strains of E. coli and genetically combined the best qualities to reach a high yield. "Other scientists thought we couldn't come close to a maximum yield," San recalled. "They said E. coli only needs to build enough lipid (fat) for its membrane and would stop."
That, as it turned out, was not true. "In fact, one of the strains we developed is very interesting: Instead of excreting the fatty acid, it wants to keep it inside. So more than 70 percent of the weight of these cells is fatty acid. These are obese E. coli," San said.
Since the project began, the researchers have increased their production 100-fold, San said. "We started with a titer of 0.4 grams per liter, and we were excited when we first produced 1 gram. Now we're up to 14 grams per liter and looking at ways to fine-tune the process. But at this point, the bug will not change that much."
Still, it will take time to scale up. "We have to be sure the bug is perfected and robust enough for industry," San said. "Strains that behave well in the lab may not do as well in an industrial setting." He said the development path will involve testing by independent labs to make sure the process is repeatable, and then initial scaling by a pilot plant in two or three years.
"I think this is a very rich area," San said. "When we started this project four years ago, nobody had paid attention to fatty acids. But I knew this would be a good model system with endless variations that could lead to real products."
More information: USDA announcement: tinyurl.com/bl3yn9n
Provided by Rice University