Novel microscopy technique tracks motion of thousands of cells at a time (w/ Video)
Researchers have developed a new way to observe and track large numbers of rapidly moving objects under a microscope, capturing precise motion paths in three dimensions.
Over the course of the study—reported online Sept. 17, 2012, in the Proceedings of the National Academy of Sciences—researchers followed an unprecedented 24,000 rapidly moving cells over wide fields of view and through large sample volumes, recording each cell's path for as long as 20 seconds.
"We can very precisely track the motion of small things, more than a thousand of them at the same time, in parallel," says research lead and National Science Foundation CAREER awardee Aydogan Ozcan, an electrical engineering and bioengineering professor at UCLA. "We were able to achieve sub-micron accuracy over a large volume, allowing us to understand, statistically, how thousands of objects move in different ways."
The latest study is an extension of several years of NSF-supported work by Ozcan and his colleagues to develop lens-free, holographic microscopy techniques with applications for field-based detection of blood-borne diseases and other areas of tele-medicine. Those efforts recently resulted in a Popular Mechanics Breakthrough Award and a National Geographic Emerging Explorer Award, among others. Ozcan's research is also supported through an NIH Director's New Innovator Award, Office of Naval Research Young Investigator Award and an Army Research Office Young Investigator Award from the Department of Defense.
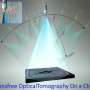
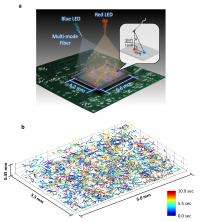
For the recent work, Ozcan and his colleagues—Ting-Wei Su, also of UCLA, and Liang Xue, of both UCLA and Nanjing University of Science and Technology in China—used offset beams of red and blue light to create holographic information that, when processed using sophisticated software, accurately reveal the paths of objects moving under a microscope. The researchers tracked several cohorts of more than 1,500 human male gamete cells over a relatively wide field of view (more than 17 square millimeters) and large sample volume (up to 17 cubic millimeters) over several seconds.
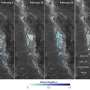
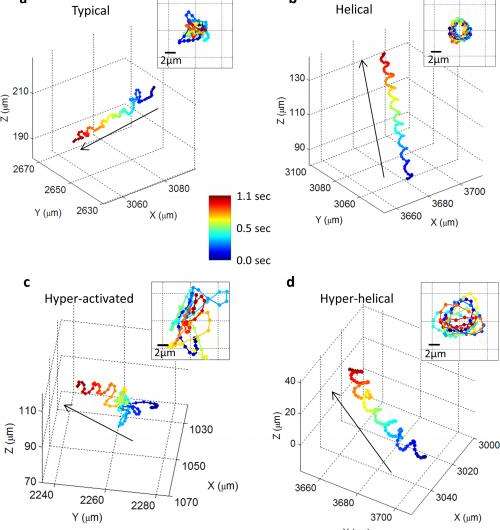
The technique, along with a novel software algorithm that the team developed to process observational data, revealed previously unknown statistical pathways for the cells. The researchers found that human male gamete cells travel in a series of twists and turns along a constantly changing path that occasionally follows a tight helix - a spiral that, 90 percent of the time, is in a clockwise (right-handed) direction.
Because only four to five percent of the cells in a given sample traveled in a helical path at any given time, researchers would not have been able to observe the rare behavior without the new high-throughput microscopy technique.
"This latest study is an extension of truly novel and creative work," says Leon Esterowitz, the NSF biophotonics program officer who has supported Ozcan's efforts. "The holographic technique could accelerate drug discovery and prove valuable for monitoring pharmaceutical treatments of dangerous microbial diseases."
The PNAS paper reports observations of 24,000 cells over the duration of the experiments. Such a large number of observations provide a statistically significant dataset and a useful methodology for potentially studying a range of subjects, from the impact of pharmaceuticals and other substances on large numbers of cells—in real time—to fertility treatments and drug development.
The same approach may also enable scientists to study quick-moving, single-celled microorganisms. Many of the dangerous protozoa found in unsanitary drinking water and rural bodies of water have only been observed in small samples moving through an area that is roughly two dimensional. The new lens-free holographic imaging technique could potentially reveal unknown elements of protozoan behavior and allow real-time testing of novel drug treatments to combat some of the most deadly forms of those microbes.
More information: High-throughput lensfree 3D tracking of human sperms reveals rare statistics of helical trajectories. PNAS, 2012 DOI: 10.1073/pnas.1212506109 . www.pnas.org/content/early/201 … /1212506109.abstract
Journal information: Proceedings of the National Academy of Sciences
Provided by National Science Foundation